Enzymatic vs. Non-Enzymatic Neuronal Detachment: A Comprehensive Guide for Cell Culture and Translational Research
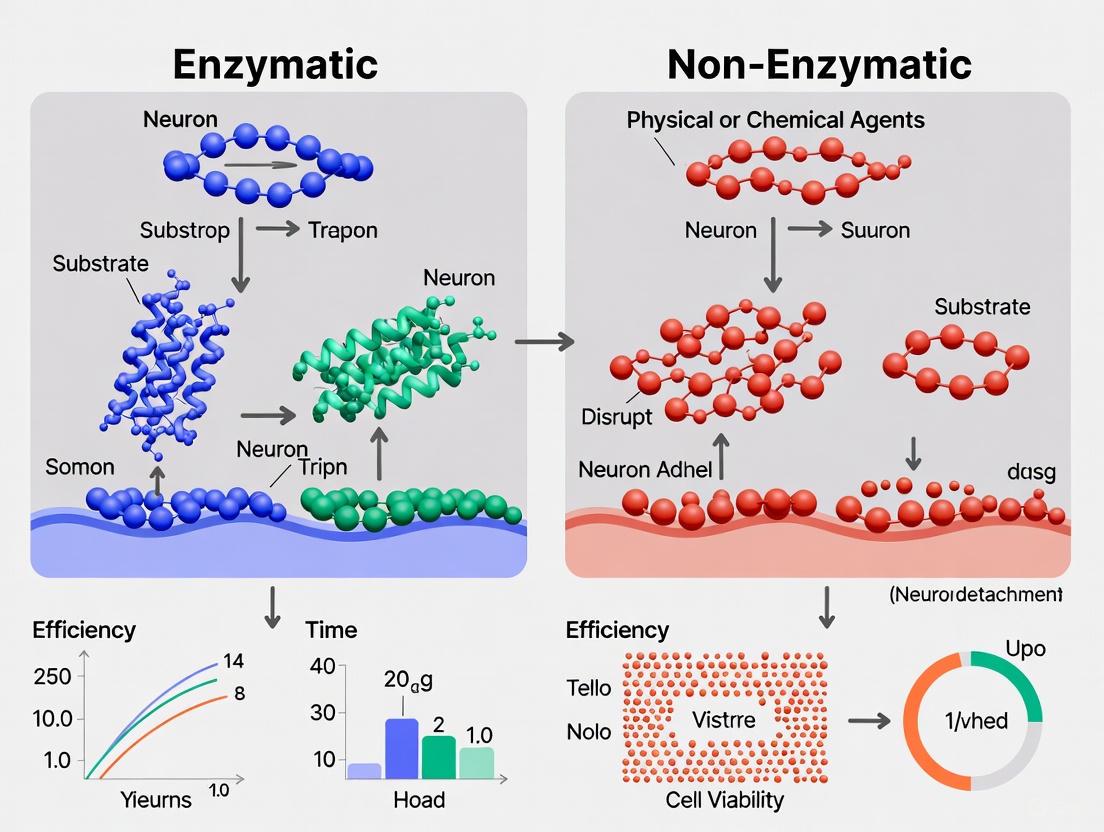
This article provides a critical comparison of enzymatic and non-enzymatic cell detachment methods, with a specific focus on applications for primary neurons and neural cultures.
Enzymatic vs. Non-Enzymatic Neuronal Detachment: A Comprehensive Guide for Cell Culture and Translational Research
Abstract
This article provides a critical comparison of enzymatic and non-enzymatic cell detachment methods, with a specific focus on applications for primary neurons and neural cultures. It explores the fundamental mechanisms of cell adhesion and the molecular-level impacts of different detachment agents on neuronal viability, surface receptors, and functionality. We detail optimized, tissue-specific protocols for dissociating sensitive neural tissues and present empirical data on cell health, yield, and phenotypic preservation. Aimed at researchers, scientists, and drug development professionals, this review synthesizes current evidence to guide method selection, troubleshoot common issues, and illuminate future directions in biomanufacturing for neuroscience and regenerative medicine.
The Building Blocks of Neuronal Adhesion: Why Your Detachment Method Matters
Understanding the Neuronal Extracellular Matrix (ECM) and Adhesion Complexes
The neuronal extracellular matrix (ECM) is a dynamic, three-dimensional network that provides not only structural support but also crucial biochemical and mechanical cues essential for neural development and function [1] [2]. Composed of macromolecules including collagens, glycosaminoglycans, elastin, proteoglycans, and specialized glycoproteins, the neuronal ECM regulates fundamental processes such as neural stem cell differentiation, neuronal migration, axonal pathfinding, and synaptic maturation [2]. The mechanical properties of the ECM, including stiffness, topology, and viscoelasticity, serve as key regulators of cellular behavior through mechanotransduction pathways, with dysregulation implicated in various neurological pathologies [1].
Cell-ECM adhesive interactions are mediated primarily by integrin receptors, which are heterodimeric proteins composed of α and β subunits that bind to ECM ligands [3]. Following binding to ECM proteins, integrins cluster together to form focal adhesion (FA) complexes, which contain structural proteins that link the ECM to the cytoskeleton and signaling effectors that regulate cell proliferation, migration, and differentiation [3]. The importance of cell-ECM adhesion is underscored by the early stage lethality in mice with genetic deletions or mutations for adhesion receptors, ligands, or associated components [3].
In neuronal contexts, specific ECM components play specialized roles. Laminins in the basal lamina are crucial for neocortical development, promoting the expansion, migration, and differentiation of neural stem cells (NSCs) [2]. Proteoglycans such as tenascin-C (Tnc) and tenascin-R (Tnr) are prominently expressed in the nervous system, with Tnc facilitating the switch from production of neuronal to glial progenitors, and Tnr inhibiting migration of NSC-derived neurons [2]. The interaction between ECM components and their receptors, particularly β1-containing integrins, constitutes the largest integrin subfamily and plays a significant role in NSC proliferation, neuronal migration, and connectivity [4].
Core Mechanisms of Cell-ECM Adhesion
Molecular Composition of Adhesion Complexes
Focal adhesions are sophisticated macromolecular assemblies that mechanically link the extracellular matrix to the intracellular actin cytoskeleton. The core molecular components include:
- Integrin receptors: Transmembrane heterodimers (α and β subunits) that directly bind ECM ligands such as fibronectin, laminin, and collagen [3]. In neural tissues, β1-containing integrins are particularly important for neural stem cell function [4].
- Talin and vinculin: Cytoskeletal adaptor proteins that connect integrins to actin filaments and recruit additional signaling molecules [3].
- Focal adhesion kinase (FAK): A key signaling mediator that transduces integrin-mediated signals to downstream pathways regulating cell survival, proliferation, and migration [3].
- α-Actinin and zyxin: Additional structural components that stabilize the adhesion complex and facilitate force transmission [3].
In the nervous system, specialized ECM structures called fractones are found in the postnatal subventricular zone (SVZ), extending tendrils of ECM between the ventricle surface to neural rosettes containing NSCs [4]. Fractones are composed of basement membrane proteins including collagen IV and various laminins, along with unique heparan sulfate and chondroitin sulfate chains that regulate NSC function [4].
Signaling Pathways in Neuronal Adhesion
Table 1: Key Signaling Pathways in Neuronal ECM Adhesion
| Pathway Component | Function in Neuronal Cells | Experimental Evidence |
|---|---|---|
| Integrin β1 | Neural stem cell proliferation, neuronal migration, connectivity [4] | Conditional knockout in mice causes abnormal neocortical lamination and cerebellar folia fusion [2] |
| YAP/TAZ | Mechanotransduction effectors regulated by ECM stiffness [1] | Activated by stiffened ECM in cancer models; promotes proliferation and survival [1] |
| Piezo1 | Mechanosensitive ion channel responding to ECM mechanical properties [1] | Transduces mechanical signals into calcium influx and downstream signaling [1] |
| TRPV4 | Mechanosensitive cation channel [1] | Responds to ECM viscoelasticity and osmotic changes [1] |
| Reelin-α3β1 integrin | Neuronal migration and cortical lamination [4] | Guides "inside-out" pattern of cortical growth; inhibition disrupts migration [4] |
The following diagram illustrates the core signaling pathways through which ECM adhesion influences neuronal behavior:
Neuronal ECM Signaling PathwaysCore mechanisms through which extracellular matrix cues influence neuronal cell behavior.
Detachment Methods: Principles and Applications
Enzymatic Detachment Methods
Enzymatic detachment utilizes proteolytic enzymes to cleave specific protein domains that mediate cell-ECM and cell-cell adhesions. The most commonly used enzymes in neuronal research include:
- Trypsin: A serine protease that cleaves peptide chains mainly at the carboxyl side of lysine or arginine residues, effectively disrupting integrin-ECM bonds and cadherin-mediated cell-cell junctions. Trypsin is widely used for general subculturing of neuronal cell lines but can damage surface receptors if overused [3].
- Accutase: A proprietary blend of proteolytic and collagenolytic enzymes that provides gentler detachment than trypsin, better preserving surface markers and cellular viability. Particularly useful for sensitive neuronal cultures and stem cell populations [5].
- Collagenase: Specifically targets collagen networks in the ECM, making it valuable for dissociating tissues with abundant collagen such as peripheral nerves or for isolating neurons from mature ECM-rich environments [3].
- Dispase: A neutral protease from Bacillus polymyxa that cleaves fibronectin, collagen IV, and to some extent collagen I, while being gentler on cell surface proteins and receptors. Often preferred for neural stem cell cultures where marker preservation is critical [2].
The efficacy of enzymatic detachment depends on multiple factors including enzyme concentration, exposure time, temperature, and the specific ECM composition of the neuronal culture system. For example, mature neuronal networks with extensive ECM deposition may require longer exposure or combination enzyme approaches [3].
Non-enzymatic Detachment Methods
Non-enzymatic approaches utilize mechanical force or chemical disruption of cell-ECM interactions without proteolytic activity:
- Chelator-based methods: EDTA and EGTA work by chelating calcium and magnesium ions that are essential for integrin-ECM binding. This approach preserves surface proteins but may not efficiently dissociate tissues or cultures with abundant ECM deposition [3].
- Mechanical detachment: Physical methods including scraping, pipetting, or hydrodynamic shear forces. These approaches can be effective but risk causing significant cellular damage and are generally unsuitable for delicate neuronal cultures or when single-cell suspensions are required [3].
- Temperature reduction: Brief cold shock treatments can weaken cell-ECM interactions by affecting membrane fluidity and receptor dynamics, though this method alone is rarely sufficient for complete detachment [3].
Recent advancements in biomaterials have led to the development of thermoresponsive surfaces that allow controlled cell detachment through temperature modulation, though these are not yet widely adopted in neuronal research contexts [6].
Comparative Analysis of Detachment Methodologies
Quantitative Comparison of Detachment Efficiency
Table 2: Efficiency Metrics of Enzymatic vs. Non-enzymatic Detachment Methods in Neuronal Cultures
| Method | Viability Recovery | Adhesion Molecule Preservation | Neurite Regrowth Capacity | Time to Detachment | Applicability to 3D Cultures |
|---|---|---|---|---|---|
| Trypsin | 70-85% [3] | Low (cleaves surface proteins) [3] | Moderate (requires re-expression) [5] | 5-15 minutes [3] | Limited (poor penetration) [3] |
| Accutase | 85-95% [5] | Moderate (partial preservation) [5] | Good [5] | 10-20 minutes [5] | Moderate [5] |
| Collagenase | 80-90% [3] | High (specific to ECM) [3] | Good [3] | 20-45 minutes [3] | Good (effective in 3D) [3] |
| EDTA/EGTA | 90-98% [3] | High (no proteolysis) [3] | Excellent [3] | 15-30 minutes [3] | Poor (surface only) [3] |
| Mechanical | 50-70% [3] | Variable (physical damage) [3] | Poor (cytoskeletal damage) [3] | Immediate [3] | Moderate (tissue fragmentation) [3] |
Functional Consequences on Neuronal Phenotype
The choice of detachment method has significant implications for downstream neuronal function and experimental outcomes:
- Neurite regeneration: Enzymatic methods requiring 24-48 hours for full neurite reextension, while chelator-based methods show more rapid recovery (4-12 hours) due to better preservation of adhesion machinery [3].
- Synaptic function: Enzymatic detachment with trypsin significantly reduces surface expression of neurotransmitter receptors and synaptic adhesion molecules, potentially altering electrophysiological properties for 3-7 days post-detachment [5].
- Gene expression profiles: Transcriptomic analyses reveal that enzymatic methods trigger stress response pathways and transient downregulation of adhesion-related genes, while mechanical methods show more variable impacts on immediate early gene expression [3].
- Stem cell differentiation: For neural stem cells, gentle enzymatic methods (Accutase) or chelator-based approaches better maintain differentiation potential compared to trypsin, which can bias lineage commitment through unintended protease-activated receptor signaling [5].
The following workflow diagram illustrates a typical experimental approach for comparing detachment methods in neuronal research:
Detachment Method Comparison WorkflowExperimental approach for evaluating enzymatic versus non-enzymatic detachment methods.
Experimental Protocols for Adhesion and Detachment Studies
Standardized Detachment Efficiency Assay
Purpose: To quantitatively compare the efficiency and cellular impact of different detachment methods on neuronal cultures.
Materials:
- Neuronal culture (primary neurons, neuronal cell lines, or neural stem cells)
- Test detachment solutions (enzymatic and non-enzymatic)
- Centrifuge capable of 300 × g
- Hemocytometer or automated cell counter
- Viability staining solution (e.g., trypan blue)
- Pre-coated culture plates for replating
Procedure:
- Culture neuronal cells under standard conditions until 70-80% confluency or desired maturity.
- Wash cells twice with appropriate buffer (PBS or Hanks' Balanced Salt Solution).
- Apply detachment solutions according to Table 2 concentrations and incubate at 37°C for recommended times.
- Gently dislodge cells by tapping or pipetting and transfer to collection tubes containing serum-containing medium to inactivate enzymes.
- Centrifuge at 300 × g for 5 minutes and resuspend in fresh medium.
- Quantify total cell count and viability using trypan blue exclusion.
- Plate equal numbers of cells into new culture vessels pre-coated with appropriate ECM substrate.
- Assess attachment efficiency after 4 hours by counting adherent cells.
- Monitor neurite outgrowth and morphological recovery at 24, 48, and 72 hours post-plating.
Data Analysis:
- Calculate detachment efficiency as (number of cells in suspension / total initial cells) × 100
- Determine viability as (viable cells / total cells) × 100
- Quantify attachment efficiency as (adherent cells after 4h / cells plated) × 100
- Measure neurite length and branching at defined timepoints using image analysis software [3]
Adhesion Strength Quantification Using Spinning Disk Assay
Purpose: To measure the force required to detach cells from ECM substrates, providing quantitative data on adhesion strength.
Materials:
- Spinning disk device [3]
- Circular coverslips coated with ECM substrates of interest
- Parallel plate flow chamber or radial flow chamber
- High-speed camera for monitoring cell detachment
- Environmental chamber to maintain 37°C and 5% CO₂ during experimentation
Procedure:
- Coat circular coverslips with defined ECM substrates (laminin, fibronectin, collagen IV, or poly-D-lysine as control).
- Seed neuronal cells at defined density and culture for desired adhesion time (typically 2-24 hours).
- Assemble spinning disk apparatus with cell-coated coverslip.
- Fill chamber with appropriate physiological buffer, potentially with viscosity enhancers like dextran to increase applied detachment force while maintaining low rotation speeds [3].
- Spin at fixed speeds for defined durations, applying known shear stresses that vary linearly with radial distance.
- Capture images at different radial distances, each corresponding to known shear stress values.
- Count adherent cells before and after spinning at each radial position.
- Calculate fraction of adherent cells versus applied shear stress.
Data Analysis:
- Generate nonlinear curve of adherent cell fraction versus applied shear stress
- Determine adhesion strength (τ₅₀) as the shear stress producing 50% cell detachment [3]
- Compare τ₅₀ values across different ECM substrates and culture conditions
- Correlate adhesion strength with FA size and composition using immunofluorescence [3]
Research Reagent Solutions for Neuronal Adhesion Studies
Table 3: Essential Reagents for Neuronal ECM and Adhesion Research
| Reagent Category | Specific Examples | Research Applications | Key Considerations |
|---|---|---|---|
| ECM Substrates | Laminin, Fibronectin, Collagen IV, Poly-D-Lysine, Chitosan [5] | Coating culture surfaces to promote specific neuronal adhesion and differentiation | Chitosan shows promise as alternative to Matrigel for supporting neuronal network development [5] |
| Proteolytic Enzymes | Trypsin, Accutase, Collagenase, Dispase [3] | Cell dissociation and subculturing; ECM degradation studies | Specificity, concentration, and exposure time critically impact surface receptor preservation [3] |
| Adhesion Inhibitors | EDTA/EGTA, RGD peptides, function-blocking integrin antibodies [3] | Studying specific adhesion mechanisms; controlled detachment | RGD peptides competitively inhibit integrin binding to fibronectin and other RGD-containing ECM proteins [3] |
| Decellularized ECM | Brain region-specific decellularized ECM (cortex, cerebellum) [7] | Providing tissue-specific ECM environments for specialized neuronal cultures | Retains tissue-specific biochemical composition and mechanical properties [7] |
| Integrin Activation Reagents | Mn²⁺, function-activating antibodies [3] | Studying inside-out activation of integrin receptors | Mn²⁺ induces constitutive integrin activation by binding to specific sites in the integrin extracellular domain [3] |
The selection between enzymatic and non-enzymatic detachment methods represents a critical methodological consideration in neuronal research, with significant implications for experimental outcomes and data interpretation. Enzymatic methods offer efficient dissociation but can compromise surface receptor integrity and alter subsequent neuronal function, while non-enzymatic approaches better preserve surface molecules but may be insufficient for robust dissociation of mature neuronal networks.
The expanding toolkit of ECM-mimetic biomaterials, including region-specific decellularized brain ECM [7] and functionalized hydrogels [6], provides new opportunities for creating more physiologically relevant neuronal culture systems. Similarly, advanced adhesion measurement technologies such as the spinning disk assay [3] enable quantitative assessment of cell-ECM interactions under controlled mechanical conditions.
Future directions in neuronal adhesion research will likely focus on developing more selective detachment strategies that target specific adhesion complexes while preserving others, allowing researchers to precisely interrogate particular molecular interactions. Additionally, the integration of real-time monitoring during detachment procedures could provide valuable insights into the dynamics of adhesion complex disassembly and inform optimized protocols for specific neuronal subtypes and experimental applications.
For researchers working with adherent cell cultures, particularly the sensitive and post-mitotic neurons, the process of cell detachment is a critical step that can significantly impact experimental outcomes and cell viability. This procedure is essential for subculturing, conducting various bioassays, and applications in tissue engineering and regenerative medicine. The fundamental challenge lies in efficiently disrupting the robust bonds between the cell and its extracellular matrix (ECM) or culture surface while preserving cellular integrity and function. The two primary approaches—enzymatic and non-enzymatic dissociation—operate through distinct mechanistic pathways, each with profound implications for downstream research, especially in neuronal studies where cell surface receptors and viability are paramount. Understanding these core mechanisms is not merely a technical exercise but a prerequisite for producing reliable and reproducible data in neuroscience and drug development.
Comparative Analysis of Detachment Mechanisms
The choice between enzymatic and non-enzymatic methods involves a key trade-off between detachment efficiency and the preservation of cell surface integrity. The following table summarizes the core characteristics of these approaches.
Table 1: Fundamental Comparison of Enzymatic vs. Non-Enzymatic Detachment Methods
| Feature | Enzymatic Methods | Non-Enzymatic Methods |
|---|---|---|
| Core Mechanism | Proteolytic cleavage of cell-surface proteins and ECM components [8]. | Physical disruption of bonds or chemical interference with cell-adhesion interactions without protein cleavage [9] [8]. |
| Primary Action | Severs anchor proteins (e.g., integrins) and degrades ECM proteins like collagen and fibronectin [8]. | Uses chelating agents (e.g., EDTA) to bind calcium, disrupting calcium-dependent adhesion [8], or applies physical stimuli like electrical current [9]. |
| Impact on Viability | Can reduce viability by damaging cell membranes and essential surface proteins, potentially boosting apoptosis [8]. | Generally maintains higher cell viability (>90%) by preserving surface protein integrity [9] [8]. |
| Impact on Surface Proteins | Destroys or damages receptors, antigens, and other proteins critical for signaling and adhesion [8]. | Better preserves native cell surface architecture, which is vital for therapeutic use and signaling studies [8]. |
| Typical Applications | Routine cell culture, high-yield dissociation from complex tissues [10]. | Sensitive cells, primary neurons, cell therapy manufacturing, and downstream assays requiring intact surface markers [9] [11] [8]. |
The quantitative performance of these methods varies significantly across key metrics, as evidenced by experimental data.
Table 2: Quantitative Performance Comparison in Cell Dissociation
| Performance Metric | Enzymatic Dissociation | Non-Enzymatic Dissociation | Experimental Context |
|---|---|---|---|
| Cell Viability | Can be compromised; varies by protocol. | >90% viability maintained [9]. | Human cancer cells detached via electrochemical method [9]. |
| Detachment Efficiency | High (>95% with optimized protocols). | Up to 95% efficiency achieved [9]. | Alternating electrochemical redox-cycling [9]. |
| Cell Yield | 25.4 ± 5.41 million cells (enzymatic) vs. 3.43 ± 0.52 million (mechanical) from 12 rat embryo spinal cords [10]. | Lower yield in mechanical dissociation; newer methods (electrical) show >5x higher yield than traditional enzymatic-mechanical method for glioblastoma tissue [12]. | Comparison of enzymatic and mechanical dissociation of embryonic rat spinal cords [10]; Electric Field Facilitated Dissociation [12]. |
| Process Time | Can be slow, from minutes to hours or overnight [12]. | Rapid; as fast as 5 minutes for some electrical methods [9] [12]. | Electric Field Facilitated Dissociation [12]. |
Detailed Experimental Protocols and Workflows
To ensure reproducibility, below are detailed methodologies for key protocols cited in this guide, highlighting the application of both enzymatic and non-enzymatic principles.
Enzymatic Dissociation of Embryonic Spinal Cord Neurons
This protocol, adapted from a comparative study, is designed to obtain a high yield of highly purified primary neurons from embryonic rat spinal cords [10].
- Tissue Isolation: Ispose spinal cords from Embryonic Day 14–15 (E14-15) rat embryos.
- Enzymatic Digestion: Mince the spinal cord tissue finely and incubate in a solution of 0.25% trypsin and 0.05% DNase I in Hanks' Balanced Salt Solution (HBSS). The digestion is carried out for 30 minutes at 37°C.
- Termination of Digestion: Stop the reaction by adding Dulbecco's Modified Eagle Medium (DMEM) supplemented with 10% Fetal Bovine Serum (FBS).
- Mechanical Dissociation: Gently triturate the tissue mixture using a fire-polished Pasteur pipette to create a single-cell suspension.
- Centrifugation and Resuspension: Centrifuge the cell suspension and resuspend the resulting pellet in a complete neuronal culture medium.
- Plating: Seed the cells onto culture vessels pre-coated with an appropriate substrate like poly-L-lysine or poly-L-ornithine.
Enzyme-Free Electrochemical Cell Detachment
This novel protocol utilizes a conductive biocompatible polymer nanocomposite surface and represents a modern non-enzymatic approach [9].
- Surface Preparation: Culture anchorage-dependent cells (e.g., human osteosarcoma or ovarian cancer cells) on a specialized conductive biocompatible polymer nanocomposite surface.
- Application of Stimulus: Apply a low-frequency alternating voltage to the culture surface. The specific optimal frequency must be determined empirically for different cell types.
- Disruption of Adhesion: The alternating electrochemical current disrupts the adhesion forces at the cell-surface interface. This process is typically completed within minutes.
- Cell Harvesting: Gently harvest the detached cells from the medium. The method allows for the recovery of over 90% of cells with high viability [9].
Coating-Based Adherence for Motor Neurons
This protocol focuses on preparing surfaces for the optimal attachment and growth of induced pluripotent stem cell-derived motor neurons (iPSC-MNs), which is a critical step before any detachment can occur [11].
- Coating Preparation: Prepare two key coating solutions:
- Poly-L-ornithine/Matrigel (POM): A mixture of poly-L-ornithine and Matrigel, a complex basement membrane matrix.
- Polyethyleneimine (PEI): A synthetic polycationic polymer solution.
- Surface Coating: Apply the chosen coating solution to the culture dishes or microelectrode array (MEA) plates and incubate overnight.
- Plating and Differentiation: Plate the iPSC-derived motor neuron progenitors onto the pre-coated surfaces and differentiate them into mature motor neurons.
- Outcome: POM coating accelerates maturation, suitable for neurodevelopmental studies. PEI coating results in a more even cell distribution and reduces the variability of electrophysiological signals, making it preferable for modeling neurodegenerative diseases like ALS [11].
Signaling Pathways and Experimental Workflows
The following diagrams illustrate the logical flow of the core mechanisms and a key experimental workflow.
Diagram 1: Core detachment mechanisms and outcomes.
Diagram 2: Coating optimization workflow for neuronal research.
The Scientist's Toolkit: Key Research Reagent Solutions
Successful cell culture and dissociation require specific, high-quality reagents. The following table details essential materials used in the featured experiments.
Table 3: Essential Reagents for Cell Dissociation and Neuronal Culture
| Reagent/Material | Function | Example Application |
|---|---|---|
| Trypsin | Protease that cleaves peptide bonds, digesting cell-adhesion proteins [8] [10]. | Standard enzymatic dissociation of tissues and adherent cell lines [10]. |
| TrypLE | A recombinant fungal trypsin-like protease, often used as an animal-origin-free alternative to trypsin [13]. | Enzymatic dissociation for surface proteomics studies [13]. |
| Collagenase | Enzyme that degrades native collagen, a key component of the ECM [12] [8]. | Dissociation of fibrous or complex tissues [12]. |
| EDTA (Ethylenediaminetetraacetic acid) | Chelating agent that binds calcium and magnesium ions, disrupting cadherin-mediated cell-cell and integrin-mediated cell-ECM adhesion [8]. | Used in non-enzymatic chelate-based detachment or in combination with enzymes [8]. |
| Poly-L-Lysine / Poly-L-Ornithine | Synthetic cationic polymers that coat surfaces, enhancing the attachment of negatively charged cell membranes [11] [14] [15]. | Pre-coating culture surfaces to improve adherence of primary neurons [14] [15]. |
| Polyethyleneimine (PEI) | A polycationic polymer resistant to proteolysis, promoting strong cell attachment and even distribution [11]. | Coating MEA plates for iPSC-derived motor neurons to reduce electrophysiological signal variability [11]. |
| Matrigel | A complex, basement membrane matrix extract containing ECM proteins like laminin and growth factors [11]. | Used as a coating, often with poly-ornithine, to support complex cell differentiation and function (e.g., POM coating) [11]. |
| Conductive Polymer Nanocomposite | A specialized smart material whose properties can be changed with electrical stimuli [9]. | Serves as the culture surface for enzyme-free electrochemical cell detachment [9]. |
The decision between enzymatic and non-enzymatic detachment methods is fundamental, dictated by the specific needs of the experiment. Enzymatic methods, while powerful and capable of high yields, act as a blunt instrument, potentially compromising cell health and surface biology. Non-enzymatic strategies, particularly modern electrochemical and advanced coating approaches, offer a more refined toolkit, prioritizing the preservation of native cell states. For neuronal research, where the integrity of the cell surface is inextricably linked to physiological function, the move towards gentler, non-enzymatic methods is not just a trend but a necessary evolution. This shift is crucial for enhancing the reliability of in vitro models, improving the success of cell-based therapies, and ultimately driving more accurate and predictive neuroscience and drug discovery.
A fundamental challenge in neuroscience research is the need to harvest cells from culture surfaces or tissues for subsequent experiments, a process that must balance high efficiency with the preservation of delicate cellular structures. For neuronal cells, whose complex morphology and surface protein expression are critical to their function, this balance is particularly vital. This guide provides an objective comparison of enzymatic and non-enzymatic cell detachment methods, framing them within the context of neuronal research to inform experimental design.
Quantitative Comparison of Detachment Methods
The choice of detachment method directly impacts key outcomes for cells. The table below summarizes experimental data comparing the performance of different techniques.
Table 1: Performance Metrics of Cell Detachment Methods
| Method | Reported Cell Viability | Detachment Efficiency/Time | Impact on Surface Proteins | Key Advantages | Key Limitations |
|---|---|---|---|---|---|
| Trypsin (Enzymatic) | 93.2% (MSC) [16] | ~5-6 min (MSC monolayers) [16] | Degrades surface proteins; can cleave FasL/Fas [8] [17] | Robust, effective, and fast for monolayers [16] | Broad proteolytic activity damages membrane integrity [8] |
| Enzyme-Free Dissociation Buffer | 68.7% (MSC) [16] | ~15-16 min (MSC monolayers) [16] | Gentler on many surface proteins [16] | Preserves structural integrity of membrane proteins [16] | Lower cell viability and reattachment rates [16] |
| Accutase (Enzymatic) | Maintains viability better than EDTA after 60 min [17] | Manufacturer's protocol: 10 min to 1 hour [17] | Cleaves specific proteins (e.g., FasL, Fas); requires ~20h recovery [17] | Considered a mild-acting enzyme for many markers [17] | Compromises specific surface proteins (FasL/Fas) [17] |
| MIT Electrochemical (Non-Enzymatic) | >90% [9] | Detachment within minutes [9] | Preserves delicate cell membranes and surface proteins [9] | High viability; automated workflow potential [9] | Emerging technology; requires specialized conductive surfaces [9] |
| Hypersonic Levitation (HLS) | 92.3% (renal cancer tissue) [18] | 15 minutes (90% utilization) [18] | Not specified; high viability suggests good preservation | High throughput, preserves rare cell populations [18] | Specialized equipment; limited data on neuronal cells [18] |
Detailed Experimental Protocols
To ensure reproducibility and critical evaluation, the methodologies from key cited studies are detailed below.
This study provides a direct, quantitative comparison of two common methods.
- Cell Culture: Human bone marrow-derived Mesenchymal Stem Cells (MSCs) were cultured in monolayers within 12-well dishes until confluent.
- Detachment Process:
- Monolayers were washed twice with calcium-free PBS.
- One milliliter of pre-warmed (37°C) 0.05% Trypsin-EDTA or enzyme-free, PBS-based dissociation buffer was added per well.
- The plates were placed in an incubator and gently pipetted every 2-3 minutes.
- Dissociation was typically complete in 5-6 minutes for trypsin and 15-16 minutes for the enzyme-free buffer.
- Assessment of Outcomes:
- Viability: The dissociated cell suspension was centrifuged, reconstituted, and analyzed using the trypan blue exclusion assay on an automated cell counter.
- Reattachment & Metabolic Activity: Dissociated cells were re-seeded onto new dishes. After 24 hours, the MTT assay was performed on the reattached cells to measure metabolic activity.
This protocol highlights the method-specific impact on surface markers, which is crucial for flow cytometry and functional studies.
- Cell Culture: RAW264.7 murine macrophage cells were used as a model for strongly adherent immune-related cells.
- Detachment Process:
- Cells were treated with either an EDTA-based non-enzymatic solution (Versene) or Accutase.
- Treatment was performed according to the manufacturer's instructions, typically involving incubation at 37°C for 10 to 30 minutes.
- Assessment of Outcomes:
- Surface Protein Levels: Detached cells were immediately analyzed by flow cytometry to measure the Mean Fluorescence Intensity (MFI) of surface markers like FasL and Fas receptor.
- Protein Cleavage: Cell lysates and supernatants from detached cells were collected for western blot analysis using an antibody against the extracellular portion of FasL to detect cleavage.
- Recovery Time: Cells detached with Accutase were re-cultured, and surface protein levels were re-assessed by flow cytometry over 20 hours to monitor recovery.
Pathways and Workflows in Cell Detachment
The following diagrams illustrate the logical workflow for method selection and the impact of enzymatic methods on critical surface components.
Diagram 1: Experimental Workflow for Method Selection
Diagram 2: Impact of Detachment on Cell Surface
The Scientist's Toolkit: Key Research Reagents
Selecting the appropriate reagents is fundamental to a successful detachment experiment. This table catalogs essential solutions and their functions.
Table 2: Essential Reagents for Cell Detachment Protocols
| Reagent / Solution | Function / Description | Common Applications |
|---|---|---|
| Trypsin-EDTA | Protease that cleaves adhesion proteins; EDTA chelates calcium to weaken integrin-mediated adhesion. [8] [19] | Standard for dissociating robust cell monolayers (e.g., MSCs, fibroblasts). [16] |
| Accutase | A blend of proteolytic and collagenolytic enzymes considered milder than trypsin. [17] | Detachment of sensitive cells, including some stem cells and immune cells. [17] |
| Enzyme-Free Dissociation Buffer | Isotonic, PBS-based solution containing chelating agents; disrupts calcium-dependent adhesion without enzymes. [16] | When preserving surface protein integrity is a priority (e.g., for flow cytometry). [16] |
| Collagenase | Enzyme that specifically breaks down native collagen, a major component of the ECM. [8] [19] | Essential for dissociating tissues rich in connective tissue, such as nerves, heart, and bone. [19] |
| Papain | A highly efficient cysteine protease that degrades myofibrillar and collagen proteins. [19] | Particularly effective for the dissociation of neural tissue with high cell viability. [19] |
| DNase I | An endonuclease that cleaves DNA. It is often added to dissociation mixes. [19] | Prevents cell clumping caused by sticky DNA released from damaged cells during tissue dissociation. [19] |
The process of detaching adherent cells is a fundamental step in neuronal research, essential for routine subculturing, cell-based assays, and therapeutic manufacturing. The method of dissociation plays a pivotal role in experimental reproducibility and outcome, as it directly impacts critical cellular attributes. This guide provides a comparative analysis of enzymatic and non-enzymatic detachment methods, focusing on their effects on viability, yield, functionality, and phenotypic stability in neural cell research. By synthesizing current experimental data, we aim to equip researchers with the evidence needed to select the most appropriate dissociation strategy for their specific applications.
Comparative Analysis of Dissociation Methods
Cell detachment strategies primarily fall into two categories: enzymatic and non-enzymatic. Enzymatic methods use proteolytic enzymes like trypsin, Accutase, and TrypLE to cleave proteins that mediate cell adhesion. Non-enzymatic methods include chelating agents (e.g., EDTA-based buffers) that sequester divalent cations critical for adhesion, as well as novel physical and electrochemical approaches.
The table below summarizes the core characteristics of these methods:
Table 1: Overview of Common Cell Detachment Methods
| Method Type | Specific Method | Mechanism of Action | Primary Applications |
|---|---|---|---|
| Enzymatic | Trypsin-EDTA | Proteolytic cleavage of adhesion proteins | General cell culture, robust dissociation [16] [8] |
| Enzymatic | Accutase | Blend of proteolytic and collagenolytic enzymes | Sensitive cells, including neural progenitors [17] [20] |
| Enzymatic | TrypLE | Recombinant fungal-derived trypsin substitute | Xeno-free culture, sensitive cells [16] [21] |
| Non-Enzymatic | EDTA-based Buffer | Chelates Ca²⁺ and Mg²⁺ ions, disrupting integrin binding | Lightly adherent cells, surface marker preservation [16] [17] |
| Non-Enzymatic | Electrochemical | Alternating current disrupts adhesion on a conductive surface | Automated biomanufacturing, high-viability harvesting [9] [22] |
| Non-Enzymatic | Mechanical Scraping | Physical dislodgement | When chemical methods are not permissible [17] [8] |
Quantitative Comparison of Key Success Metrics
Cell Viability and Yield
Cell viability post-detachment is a primary metric for assessing method gentleness. Yield, or the number of cells recovered, is equally critical for applications requiring large cell numbers.
Table 2: Comparison of Viability and Yield Metrics
| Detachment Method | Cell Type | Viability (%) | Yield / Detachment Efficiency | Citation |
|---|---|---|---|---|
| Trypsin | Mesenchymal Stem Cells (MSC) | 93.2% ± 3.2 | High | [16] |
| Enzyme-free Buffer | Mesenchymal Stem Cells (MSC) | 68.7% ± 5.0 | Significantly lower | [16] |
| Electrochemical | Osteosarcoma & Ovarian Cancer Cells | > 90% | 95% detachment efficiency | [9] [22] |
| Accutase | Neural Progenitor Cells | High (Inferred from efficacy) | Effective for single-cell suspension | [20] |
| Mechanical Scraping | Macrophages | Preserved (Context-dependent) | High (but risks cell damage) | [17] |
Key Findings:
- Trypsin demonstrates superior viability and yield for MSCs compared to standard enzyme-free buffers [16].
- Novel electrochemical methods show great promise, reporting over 90% viability and 95% detachment efficiency, overcoming key limitations of chemical methods [9] [22].
- For neural stem and progenitor cells, which grow in tight clusters, Accutase, TrypLE, and trypsin are more effective at creating single-cell suspensions than mechanical means alone, a necessity for accurate counting and downstream assays [20].
Phenotypic Stability and Surface Marker Preservation
Preserving the native surface proteome is crucial for immunophenotyping, signaling studies, and functional assays. Different detachment methods variably affect cell surface markers.
Table 3: Impact on Cell Surface Markers and Phenotype
| Detachment Method | Cell Type | Effect on Surface Markers / Phenotype | Citation |
|---|---|---|---|
| Accutase | Macrophages | Significantly decreases surface FasL and Fas receptor; cleaves extracellular portion of FasL. | [17] |
| Accutase | Human Monocyte-Derived Macrophages | Selectively cleaves M2 markers CD206 and CD163. Effect is variable across donors. | [23] |
| EDTA-based Buffer | Macrophages | Preserves surface FasL and Fas receptor better than Accutase. | [17] |
| Scraping | Macrophages | Best preservation of surface FasL levels compared to all chemical methods. | [17] |
| Trypsin | General Cell Types | Can cleave surface proteins and receptors, dysregulating protein expression and metabolic pathways. | [8] |
Key Findings:
- Accutase, often considered a gentle enzyme, can significantly compromise specific surface proteins like FasL and Fas receptor in macrophages, and cleave polarization markers like CD206 and CD163 [17] [23].
- Non-enzymatic methods, particularly EDTA-based buffers and scraping, offer superior preservation of many surface markers [17].
- The effects of Accutase are reversible; surface protein levels can recover after approximately 20 hours in culture [17].
Cellular Functionality
The ultimate test of a detachment method is whether the harvested cells remain functional.
Endocytic Function: In human monocyte-derived macrophages, the process of enzymatic detachment itself was found to impair the cells' endocytic ability, a key macrophage function [23].
Reattachment and Proliferation: A critical metric for culture expansion is the ability of dissociated cells to reattach and proliferate. For MSCs, the proportion of viable cells that reattach 24 hours after dissociation is significantly lower for cells obtained with enzyme-free buffer compared to trypsin. This trend holds true even after a freeze-thaw cycle [16].
Detailed Experimental Protocols
To ensure reproducibility, below are detailed methodologies from key cited studies.
Objective: To compare the effectiveness of trypsin and enzyme-free dissociation buffer in harvesting viable MSCs with high reattachment potential.
Materials:
- Confluent monolayers of MSC in 12-well plates
- 0.05% (w/v) Trypsin-EDTA solution, pre-warmed to 37°C
- Enzyme-free, PBS-based cell dissociation buffer, pre-warmed to 37°C
- Ca²⁺-free Phosphate Buffered Saline (PBS)
- Cell culture medium (e.g., MSCGM bullet kit)
Procedure:
- Wash confluent MSC monolayers twice with Ca²⁺-free PBS.
- Add 1 ml of pre-warmed trypsin or enzyme-free dissociation buffer to each well.
- Place the culture dish in a 37°C incubator and gently pipet the solution every 2-3 minutes.
- Average incubation: 5-6 min for trypsin; 15-16 min for enzyme-free buffer.
- Once cells are detached, collect the cell suspension in a microcentrifuge tube.
- Centrifuge at 500×g for 5 minutes. Discard the supernatant.
- Resuspend the cell pellet in 0.5 ml PBS (with Ca²⁺) for immediate viability analysis OR in 1.0 ml culture medium for re-seeding.
Assessment:
- Viability: Analyze using trypan blue exclusion assay with an automated cell counter.
- Reattachment: Re-seed dissociated cells at a known density into new dishes. After 24 hours, wash off unattached cells and perform an MTT assay on the reattached cells.
Objective: To evaluate the impact of different detachment methods on the surface expression of Fas Ligand (FasL) and Fas receptor.
Materials:
- Adherent cell cultures (e.g., RAW264.7 macrophages)
- Accutase solution
- EDTA-based non-enzymatic detachment solution (e.g., Versene)
- Cell scrapers
- Complete cell culture medium
- Antibodies for flow cytometry (e.g., against FasL, Fas receptor, F4/80)
Procedure:
- Culture cells to appropriate confluence.
- For each detachment method:
- Accutase/EDTA: Aspirate medium, add detachment solution, and incubate for 10-30 minutes at 37°C. Gently tap or pipet to detach.
- Scraping: Aspirate medium, add a small volume of buffer, and physically dislodge cells using a cell scraper.
- Collect cells from all methods and wash with complete medium.
- For recovery experiments, seed the harvested cells in fresh complete medium and analyze at various time points (e.g., 2, 6, 20 hours post-detachment).
- Analyze surface marker expression via flow cytometry immediately after detachment or after recovery.
The Scientist's Toolkit: Essential Research Reagents
Table 4: Key Reagents for Cell Detachment Studies
| Reagent / Solution | Function in Research | Key Considerations |
|---|---|---|
| Trypsin-EDTA | Gold-standard enzymatic dissociation. | Robust but may damage sensitive surface proteins; animal-derived. [16] [8] |
| Accutase | Gentle enzymatic dissociation for delicate cells. | Effective for neural clusters; but can cleave specific markers (FasL, CD163). [17] [20] |
| TrypLE Express | Recombinant, xeno-free alternative to trypsin. | Consistent formulation, suitable for therapeutic manufacturing. [16] [21] |
| EDTA-based Buffer | Non-enzymatic dissociation via chelation. | Preserves surface proteins; may be insufficient for strongly adherent cells. [16] [17] |
| Cell Dissociation Scraper | Mechanical detachment. | Bypasses chemical effects; risk of shear stress and cell lysis. [17] |
| MTT Reagent | Assesses metabolic activity of reattached cells. | Proxy for post-detachment viability and health. [16] |
| Trypan Blue | Dye exclusion test for immediate cell viability. | Standard, quick assessment post-detachment. [16] [20] |
Decision Framework and Visual Workflows
Selecting the optimal detachment method requires balancing your research goals with the known impacts of each technique. The following diagram illustrates the key decision-making pathway and the underlying molecular mechanisms affected by different methods.
Diagram 1: Method Selection Workflow
The biochemical and physical mechanisms of action for detachment methods directly influence cellular outcomes. The following diagram summarizes how different methods interact with cell adhesion structures.
Diagram 2: Mechanisms of Cell Detachment Methods
The choice between enzymatic and non-enzymatic cell detachment methods is not a one-size-fits-all decision but a strategic one based on the specific requirements of the experiment. Enzymatic methods like trypsin and Accutase generally offer robust dissociation and high viability for many cell types, including challenging-to-dissociate neural progenitor clusters [20]. However, this efficiency comes at the cost of altering the cell surfaceome, potentially cleaving critical receptors and markers, which can confound downstream phenotypic and functional analyses [17] [23]. Traditional non-enzymatic methods, such as EDTA buffers and scraping, excel at preserving surface marker integrity but may lag in detachment efficiency and can compromise yield or viability [16] [17].
Emerging technologies, particularly electrochemical detachment, present a compelling future direction. This enzyme-free strategy reports detachment efficiencies of 95% with viabilities exceeding 90%, addressing key limitations of both traditional enzymatic and non-enzymatic methods [9] [22]. Such advances highlight a growing trend toward integrating physical principles with material science to create gentler, more controllable, and automatable cell harvesting solutions for advanced applications in regenerative medicine and large-scale biomanufacturing [24] [8] [25].
In conclusion, researchers must weigh the trade-offs between dissociation efficiency, cell viability, and the preservation of phenotypic and functional integrity. By aligning the detachment method with the primary research metric of importance, scientists can ensure the reliability and reproducibility of their work in neuronal research and beyond.
Practical Protocols: Implementing Detachment Techniques in Neural Cell Culture
The quest to obtain viable, intact single cells from neuronal and brain tissues is a fundamental prerequisite for advanced research in neuroscience, drug discovery, and cell therapy. The dissociation process must carefully balance efficiency with the preservation of cell viability, surface markers, and physiological function. Enzymatic methods remain the cornerstone of this process, with trypsin, collagenase, and Accutase emerging as the most prominent workhorses. Each enzyme offers a distinct mechanism of action, leading to variations in cell yield, viability, and suitability for specific downstream applications. This guide objectively compares these three enzymatic agents, framing the analysis within the broader scientific discussion of enzymatic versus non-enzymatic detachment methods. We present summarized experimental data, detailed methodologies from key studies, and practical tools to assist researchers, scientists, and drug development professionals in selecting the optimal dissociation strategy for their experimental needs.
Comparative Analysis of Enzymatic Agents
The following table synthesizes data from multiple studies to provide a direct comparison of the three primary enzymatic agents, highlighting their key characteristics and experimentally observed outcomes.
Table 1: Direct Comparison of Trypsin, Collagenase, and Accutase for Cell Dissociation
| Feature | Trypsin | Collagenase | Accutase |
|---|---|---|---|
| Origin & Composition | Serine protease, often animal-derived (porcine/porcine) [26] [27] | Metalloprotease from Clostridium histolyticum; mixture of collagenolytic enzymes [26] [28] | Defined blend of proteolytic and collagenolytic enzymes; non-animal origin [27] [29] |
| Primary Mechanism | Cleaves peptide bonds, particularly at lysine and arginine residues [27] | Hydrolyzes native triple-helical collagen in the extracellular matrix (ECM) [30] [28] | Combined proteolytic and collagenolytic activity targeting multiple ECM components [27] |
| Typical Cell Viability | ~93% (MSC monolayers) [16] | Varies by type; Collagenase D preserves surface proteins [30] | ~75% (microglia from adult mouse brain); superior to trypsin in neural stem cells (90-95% vs 70-80%) [26] [29] |
| Impact on Surface Markers | Can damage surface epitopes and receptors; cleaves surface proteins [26] [30] | Collagenase D recommended for surface protein integrity [30] | Better preservation of surface markers for flow cytometry [27] [29] |
| GMP/Clinical Suitability | Concerns due to animal origin; not ideal for ATMPs [26] | Used in clinical cell therapy production [31] | GMP-conform, non-animal origin makes it suitable for ATMPs [26] |
| Key Advantages | Highly efficient, fast-acting, low cost [16] [27] | Effective on dense tissues rich in collagen; gentler on some surface proteins [30] [28] | Gentle, ready-to-use, no inactivation required, preserves epitopes [27] |
| Key Disadvantages | Harsh, animal-origin, requires inactivation, can harm surface proteins [26] [27] | Slower on non-collagenous ECM, can be less specific [28] | Slayer dissociation time for some cell types, may be less effective on very dense tissues [16] |
Experimental Data and Performance
Controlled studies provide critical quantitative data for evidence-based protocol selection. The following tables consolidate key findings on cell viability and dissociation efficiency across different tissue and cell models.
Table 2: Experimental Cell Viability Outcomes from Comparative Studies
| Cell / Tissue Type | Trypsin Viability | Collagenase Viability | Accutase Viability | Study Context |
|---|---|---|---|---|
| Mesenchymal Stem Cells (MSC) | 93.2% [16] | Information missing | 68.7% [16] | Dissociation of adherent monolayers [16] |
| Microglia (Mouse Brain) | Data missing | Data missing | 75% (highest yield, low variance) [29] | Flow cytometry preparation; outperformed dispase, papain, trypsin [29] |
| Neural Stem Cells | 70-80% [26] | Information missing | 90-95% [26] | Cell detachment and viability post-dissociation [26] |
| Human Brain Tumors & Tissues | Information missing | ~85% (with other enzymes) [28] | Information missing | Neutral Protease (NP) achieved 85-93% viability [28] |
Table 3: Dissociation Efficiency and Reattachment Metrics
| Metric | Trypsin | Enzyme-Free Buffer | Notes |
|---|---|---|---|
| Time for MSC Monolayer Dissociation | ~5-6 minutes [16] | ~15-16 minutes [16] | With gentle pipetting [16] |
| Viable Cell Reattachment Rate (24h post-dissociation) | Significantly higher [16] | Significantly lower [16] | Assessed via MTT assay [16] |
Detailed Experimental Protocols
To ensure reproducibility, this section outlines specific methodologies cited in the comparative data.
This protocol is derived from the study that generated the viability data in Table 1 and Table 3.
- Step 1: Preparation. Confluent human MSC monolayers cultured in 12-well dishes are washed two times with Ca²⁺-free PBS. Both 0.05% (w/v) Trypsin-EDTA and enzyme-free PBS-based dissociation buffer are pre-warmed to 37°C.
- Step 2: Enzymatic Dissociation. After washing, 1 ml of pre-warmed trypsin or enzyme-free dissociation buffer is added to each well. The plates are placed in a cell culture incubator and subjected to gentle pipetting every 2-3 minutes.
- Step 3: Monitoring and Harvesting. Dissociation is monitored. Trypsin typically requires 5-6 minutes, while the enzyme-free buffer requires 15-16 minutes. The dissociated cell suspension is collected and centrifuged at 500×g for 5 minutes.
- Step 4: Viability Assessment. The cell pellet is reconstituted in PBS (with Ca²⁺). Cell viability is assessed using the trypan blue exclusion assay with an automated cell counter.
- Step 5: Reattachment Assay (MTT). For reattachment analysis, dissociated cells are re-seeded into new culture dishes. After 24 hours, unattached cells are washed off, and the reattached viable cells are assessed using the MTT assay.
This protocol established Accutase as a superior enzyme for isolating microglia with high yield and low variance.
- Step 1: Tissue Preparation. Adult mice are transcardially perfused with cold PBS. The brains are isolated, placed in ice-cold HBSS, and mechanically minced into the smallest parts possible with a surgical scalpel.
- Step 2: Enzymatic Digestion. 2 mL of Accutase is added to each brain sample. Samples are incubated for 30 minutes at 37°C.
- Step 3: Enzyme Inactivation. After incubation, the enzyme is inactivated by adding 4 mL of DMEM.
- Step 4: Myelin Removal. The cell suspension is centrifuged, and the pellet is subjected to myelin removal using a Percoll gradient to purify the microglial population for flow cytometry analysis.
This methodology identified Neutral Protease (NP), a specific class of collagenase, as highly effective for sensitive tissues.
- Step 1: Sample Preparation. Freshly resected brain tumor (BT) or non-tumorous brain tissue is cleansed of clots and necrotic areas, weighed, and cut into 1-2 mm pieces.
- Step 2: Slurry Creation. The tissue is resuspended in HBSS (with Ca²⁺ and Mg²⁺) at a concentration of 100 mg tissue per ml.
- Step 3: Enzymatic Digestion. The slurry is divided into tubes, and Neutral Protease (NP) from Clostridium histolyticum is added at an optimal concentration of 0.11 DMC u/ml.
- Step 4: Incubation. Tubes with unlocked caps are incubated either at 37°C for 30-120 minutes or at room temperature overnight.
- Step 5: Trituration and Harvest. Following incubation, the tissue is triturated 5-8 times using a 5 ml plastic Pasteur pipette. The cell mixture is left to settle for ~30 seconds, and large undigested debris is discarded. The supernatant containing single cells is washed twice with PBS (without Ca²⁺ and Mg²⁺).
Workflow and Decision Pathway
The following diagram illustrates the logical decision-making process for selecting and implementing a cell dissociation method, integrating key considerations from the presented data.
The Scientist's Toolkit: Essential Research Reagent Solutions
Selecting the right reagents is critical for successful dissociation. The following table details key solutions used in the protocols and studies discussed in this guide.
Table 4: Key Reagents for Cell Dissociation Protocols
| Reagent / Solution | Primary Function in Dissociation | Example Use Case |
|---|---|---|
| Trypsin-EDTA | Proteolytic enzyme cleaves cell-adhesion proteins; EDTA chelates calcium/magnesium to enhance detachment [16] [27] | Rapid dissociation of robust, adherent cell monolayers (e.g., MSCs) where surface marker integrity is not the primary concern [16]. |
| Accutase | Ready-to-use blend of proteases and collagenases that acts gently on cells and preserves surface epitopes [27] [29] | Detachment of sensitive cells like neurons and stem cells; preparation of single-cell suspensions for flow cytometry [26] [29]. |
| Collagenase (Type IV/D) | Metalloprotease that hydrolyzes native collagen in the extracellular matrix, crucial for digesting structural tissue [30] [28] | Dissociation of dense tissues like solid tumors or primary organs where collagen is a major ECM component [28]. |
| Hank's Balanced Salt Solution (HBSS) | Salt solution providing an ionic and nutrient-balanced environment to maintain cell viability during processing outside the incubator [29] [28] | Washing tissue samples, creating tissue slurries, and as a base for enzyme solutions during dissociation [29]. |
| Percoll / Sucrose Solutions | Density gradient media used to separate and purify specific cell populations (e.g., microglia) from debris and myelin post-digestion [29] | Purification of microglia from total brain cell suspensions after enzymatic digestion for downstream flow cytometry [29]. |
| DNase I | Endonuclease that degrades DNA released from dead cells, reducing solution viscosity ("gooeyness") and preventing cell clumping [28] | Added to digestion mixes of tissues with high rates of cell death (e.g., tumors) to improve sample quality and cell yield [28]. |
| Fetal Bovine Serum (FBS) / Human Platelet Lysate | Contains trypsin inhibitors and proteins that inactivate trypsin; also used as a supplement in culture media [16] [31] | Stopping the reaction of trypsin digestion to prevent over-digestion and cell damage [16]. |
In neuronal research, the process of detaching adherent cells for subculturing or analysis is a fundamental yet critical step. This guide objectively compares the performance of non-enzymatic detachment methods—primarily chelating agents like EDTA and physical scraping—against enzymatic alternatives. The integrity of cell surface molecules and overall cell viability following detachment are paramount for downstream applications, from flow cytometry to functional assays. Within the broader thesis comparing enzymatic and non-enzymatic detachment for neuronal research, this article provides a data-driven evaluation to help researchers select the most appropriate method for their experimental goals.
Performance Comparison: Key Experimental Data
The following tables summarize quantitative data from studies comparing the performance of different cell detachment methods.
Table 1: Cell Viability and Reattachment Efficiency After Detachment
| Detachment Method | Cell Type | Viability Post-Detachment | Reattachment Efficiency (24h) | Source |
|---|---|---|---|---|
| Trypsin-EDTA | Mesenchymal Stem Cells (MSC) | 93.2% ± 3.2% | High (Referent) | [16] |
| Enzyme-Free Dissociation Buffer (EDTA-based) | Mesenchymal Stem Cells (MSC) | 68.7% ± 5.0% | Significantly Lower | [16] |
| Accutase | Macrophages (RAW264.7) | Maintained higher viability vs. EDTA after 60-90 min | Not Reported | [17] |
| Scraping | Macrophages (RAW264.7) | Not Reported | Preserved highest surface FasL levels | [17] |
Table 2: Impact on Cell Surface Marker Integrity
| Detachment Method | Effect on Surface Marker FasL | Effect on Surface Marker Fas Receptor | Recovery Time for Surface Proteins | Source |
|---|---|---|---|---|
| EDTA-based Buffer | Moderate decrease vs. scraping | Moderate decrease vs. scraping | Not Reported | [17] |
| Accutase | Significant decrease; cleaved into fragments | Significant decrease | ~20 hours | [17] |
| Scraping | Preserved highest levels | Not Reported | Not Applicable | [17] |
Mechanisms of Action and Experimental Workflows
How Non-Enzymatic Methods Work
Non-enzymatic detachment methods function primarily by disrupting the divalent cation bridges (particularly Ca²⁺ and Mg²⁺) that are essential for integrin-mediated cell adhesion to the extracellular matrix (ECM) and for cadherin-mediated cell-cell contacts [16] [17]. EDTA (Ethylenediaminetetraacetic acid) is a chelating agent that binds these cations with high affinity. By sequestering them, EDTA causes the integrins to lose their binding capability, leading to a weakening of cell adhesion and eventual detachment.
In the context of neuronal research, the regulation of cell-cell adhesion is particularly crucial during processes like the chain migration of neuroblasts. Studies have shown that a Fyn-mediated signaling pathway, which can be influenced by adhesion dynamics, regulates the detachment of neuroblasts from chains in the postnatal olfactory bulb by controlling N-cadherin-based adherens junctions [32]. This in vivo process shares a conceptual parallel with in vitro detachment, as both involve the precise regulation of cell adhesion.
Diagram 1: Mechanism of EDTA-Based Cell Detachment.
Detailed Experimental Protocols
Protocol 1: Detaching Adherent Cells with EDTA-based Buffer
This protocol is adapted from methods used in macrophage and stem cell studies [16] [17].
- Preparation: Pre-warm the enzyme-free, EDTA-based phosphate-buffered saline (PBS) dissociation buffer (e.g., Gibco Versene) to 37°C. Ensure the confluent cell monolayer (e.g., in a 12-well plate) is ready for harvest.
- Washing: Aspirate the culture medium. Gently wash the cell monolayer two times with Ca²⁺-free and Mg²⁺-free PBS to remove residual media and divalent cations.
- Application: Add enough pre-warmed EDTA-based dissociation buffer to cover the monolayer (e.g., 1 mL per well of a 12-well plate).
- Incubation: Place the culture vessel in a 37°C incubator for approximately 15-30 minutes. Observe cells periodically under a microscope. The cells will begin to round up and detach. For strongly adherent cells, gentle mechanical dislodgement by tapping the vessel or careful pipetting may be required.
- Neutralization & Collection: Once the majority of cells are detached, add complete cell culture media (containing serum, which helps neutralize the EDTA) to the vessel. Gently pipette the solution across the surface to collect all cells and ensure a single-cell suspension.
- Centrifugation: Transfer the cell suspension to a centrifuge tube and spin at 500 × g for 5 minutes. Discard the supernatant containing the dissociation buffer.
- Resuspension: Resuspend the cell pellet in an appropriate volume of fresh, pre-warmed culture medium for subsequent counting and re-plating.
Protocol 2: Detaching Cells by Scraping
This physical method is useful when preserving surface proteins is the highest priority [17].
- Preparation: Pre-cool PBS and culture media on ice if aiming to minimize metabolic activity post-detachment.
- Washing: Aspirate the culture medium and gently wash the monolayer with PBS.
- Scraping: Add a small volume of cold PBS or serum-free medium to keep cells moist. Using a sterile, flexible cell scraper, gently but firmly scrape the entire surface of the culture vessel. Use a consistent, linear motion to avoid foaming and excessive shear stress.
- Collection: Pipette the cell suspension containing the detached cells into a centrifuge tube. Rinse the culture surface with additional medium to collect any remaining cells.
- Centrifugation and Resuspension: Centrifuge the collected suspension at 500 × g for 5 minutes. Resuspend the cell pellet in the desired medium for downstream applications. Note that this method often results in clusters of cells rather than a perfect single-cell suspension.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Non-Enzymatic Cell Detachment
| Reagent/Material | Function/Description | Example Use Case |
|---|---|---|
| EDTA-based Dissociation Buffer | Isotonic, enzyme-free solution of salts and chelating agents in PBS. Gently disrupts integrin function. | Detaching lightly adherent cells; flow cytometry where key surface antigens are sensitive to enzymatic cleavage [16] [17]. |
| Cell Scraper | Sterile, flexible plastic blade attached to a handle for mechanically dislodging cells. | Harvesting strongly adherent cells (e.g., primary macrophages) where surface protein integrity is critical and viability is less of a concern [17]. |
| Calcium-/Magnesium-Free PBS | Phosphate-buffered saline without Ca²⁺ and Mg²⁺. Used for washing steps to pre-emptively weaken adhesion. | Essential preparatory step before adding EDTA-based detachment buffers to maximize efficiency [16]. |
| Complete Culture Medium (with Serum) | Used to neutralize the effect of EDTA after detachment, stopping the process and providing nutrients for recovery. | Resuspending cells post-detachment to prevent prolonged exposure to chelating agents [17]. |
The choice between non-enzymatic detachment methods is a trade-off. EDTA-based buffers offer a gentler, more reproducible alternative to harsh enzymes for creating single-cell suspensions, but with potentially lower viability and efficiency for some cell types. Physical scraping guarantees the preservation of sensitive surface epitopes but sacrifices single-cell suspension quality and can be more traumatic to cells. The optimal method is dictated by the specific cell type, the primary requirement of the downstream application (e.g., surface protein integrity vs. high viability), and the necessary balance between experimental convenience and biological fidelity.
The isolation and culture of primary neurons from specific regions of the rat nervous system represent a fundamental methodology for investigating neuronal function, development, and pathology in vitro [33]. These cultured neurons closely mimic the in vivo environment, providing physiologically relevant data for studying neurodegenerative disorders such as Alzheimer's and Parkinson's disease, pathological mechanisms, and therapeutic strategies [33]. The dissociation process—breaking down neural tissue into viable single-cell suspensions—serves as the critical first step that directly impacts the success of all downstream applications, from single-cell sequencing and electrophysiological studies to the establishment of reliable in vitro models of both the central and peripheral nervous systems [33] [12].
The broader thesis framing this technical discussion centers on the ongoing methodological comparison between enzymatic and non-enzymatic detachment approaches for neuronal research. While enzymatic methods have long dominated standard laboratory practice, emerging non-enzymatic technologies present distinct advantages and limitations that researchers must carefully consider based on their specific applications [12] [8]. Enzymatic dissociation typically utilizes proteases like trypsin and collagenase to digest extracellular matrix components and intercellular junctions, but these enzymes can potentially damage cell surface proteins, affect cellular metabolism, and introduce artifacts in downstream analyses [12] [8]. Conversely, non-enzymatic alternatives—including mechanical, acoustic, and electrical approaches—aim to preserve cellular integrity but may present challenges in yield, reproducibility, and standardization across different tissue types [12] [18] [8].
This guide provides a detailed, evidence-based comparison of optimized dissociation protocols for three key neural tissues: the cortex, hippocampus, and dorsal root ganglia (DRG). By presenting structured experimental data, step-by-step methodologies, and comparative analyses of cellular outcomes, we aim to equip researchers with the practical knowledge needed to select and implement the most appropriate dissociation strategy for their specific research objectives in neuroscience and drug development.
Quantitative Comparison of Dissociation Outcomes
The effectiveness of neuronal dissociation protocols is quantitatively assessed through several key metrics: cell viability, cell yield, and purity. These parameters vary significantly based on the neural tissue type, developmental stage, and specific dissociation methodology employed. The tables below summarize optimal experimental outcomes achieved through region-specific protocol optimization.
Table 1: Tissue-Specific Dissociation Parameters and Outcomes
| Neural Tissue | Developmental Stage | Optimal Dissociation Method | Reported Viability | Key Applications |
|---|---|---|---|---|
| Cortex | Embryonic Day 17-18 (E17-E18) | Enzymatic (Trypsin) + Mechanical Trituration | High viability maintained [33] | Neurodegenerative disease modeling, network studies [33] |
| Hippocampus | Postnatal Day 1-2 (P1-P2) | Enzymatic (Trypsin) + Mechanical Trituration | High viability maintained [33] | Synaptic plasticity, memory studies, toxicity testing [33] |
| Dorsal Root Ganglia (DRG) | Young Adult (6-week-old) | Enzymatic (Collagenase/Trypsin) + Mechanical Trituration | High viability maintained [33] | Pain research, peripheral neuropathy, sensory neuron function [34] |
Table 2: Comparative Analysis of General Dissociation Techniques
| Technique | Key Reagents/Equipment | Relative Viability | Relative Yield | Key Advantages | Key Limitations |
|---|---|---|---|---|---|
| Traditional Enzymatic | Trypsin, Collagenase, DNAse | Variable (Can be >90%) [12] | High | High efficiency, well-established protocols [12] | Potential damage to surface epitopes [8] |
| Non-Enzymatic Chemical | EDTA, Chelator-based solutions | Preserves surface markers [8] | Moderate | Preserves surface proteins [8] | Less effective for tough matrices [8] |
| Advanced Non-Enzymatic (HLS) | Hypersonic Levitation System | 92.3% [18] | 90% tissue utilization [18] | Non-contact, preserves rare cells [18] | Specialized equipment required [18] |
| Electrical Dissociation | Electric Field Application | ~80% [12] | >5x higher vs. traditional [12] | Rapid (5 min) [12] | Tissue-specific optimization needed [12] |
Detailed Experimental Protocols
Materials and Reagent Preparation
The Scientist's Toolkit: Essential Reagents and Equipment
Table 3: Core Reagents and Solutions
| Item | Function/Application |
|---|---|
| Neurobasal Plus Medium | Base culture medium for CNS neurons (cortex, hippocampus, spinal cord) [33] |
| F-12 Medium | Base culture medium for DRG neurons [33] |
| B-27 Supplement | Serum-free supplement essential for neuronal survival and growth [33] |
| Nerve Growth Factor (NGF) | Critical trophic factor for DRG neuron survival and maturation [33] |
| Collagenase/Trypsin | Enzymes for digesting extracellular matrix and cell-cell junctions [33] [34] |
| Hanks' Balanced Salt Solution (HBSS) | Isotonic buffer for tissue dissection and washing [33] |
| Poly-D-Lysine | Substrate for coating culture vessels to promote neuronal adhesion [33] |
Table 4: Essential Laboratory Equipment
| Equipment | Function |
|---|---|
| #5 Fine Forceps | Precise dissection of neural tissues [33] |
| CO2 Chamber | Euthanasia of donor animals [33] |
| Tissue Culture Hood | Maintains aseptic conditions for all dissociation steps [33] |
| Water Bath | Warming enzymes and culture media to optimal temperature [33] |
| Centrifuge | Pellet cells after dissociation [33] |
Coating Culture Vessels
- Prepare Coating Solution: Dilute poly-D-lysine in sterile distilled water to a concentration of 0.1 mg/mL.
- Coat Surfaces: Add sufficient solution to cover the bottom of the culture plate or flask.
- Incubate: Leave the coated vessels in a tissue culture incubator (37°C) for a minimum of 2 hours, or alternatively, overnight at room temperature.
- Rinse: Before use, thoroughly rinse the coated surfaces three times with sterile distilled water to remove any excess, unbound poly-D-lysine.
- Air Dry: Allow the vessels to air dry completely under the tissue culture hood before plating cells [33].
Region-Specific Dissociation Protocols
Embryonic Rat Cortex and Hippocampal Dissociation
Tissue Dissection:
- Animal Source: Sacrifice a timed-pregnant E17-E18 rat dam using a CO2 chamber, confirming death by lack of response to mechanical stimuli.
- Extract Embryos: Place the dam in a supine position, perform a laparotomy to expose the uterine horn, and carefully transfer the embryos to a 60-mm culture dish filled with ice-cold HBSS.
- Remove Brain: Under a dissection microscope, position the embryo prone. Use two #5 fine forceps to gently remove the skin and skull, exposing the brain. Carefully extract the whole brain and transfer it to a fresh dish with cold HBSS.
- Isolate Cortex/Hippocampus: Place the brain in a dorsal view. Carefully separate the cerebral hemispheres, removing the meninges. For the cortex, collect the cortical tissue. For the hippocampus, identify the dark C-shaped structure within the hemisphere and carefully isolate it [33].
Enzymatic Digestion:
- Mince Tissue: Use a sterile scalpel or fine scissors to mince the collected cortical or hippocampal tissue into a fine slurry.
- Digest: Transfer the minced tissue to a 15-mL tube containing a pre-warmed (37°C) enzymatic solution (e.g., 0.125-0.25% trypsin). Incubate for 15-20 minutes in a water bath, gently inverting the tube periodically [33].
Mechanical Trituration and Plating:
- Terminate Digestion: Add 1 mL of MEM-10 (Eagle's Minimal Essential Medium with 10% Fetal Bovine Serum) to neutralize the trypsin. Centrifuge the tube at 2000 rpm for 5 minutes to pellet the tissue.
- Triturate: Carefully aspirate the supernatant. Resuspend the pellet in fresh, pre-warmed neuronal culture medium (Neurobasal Plus supplemented with B-27, GlutaMAX, and Penicillin/Streptomycin). Using a fire-polished Pasteur pipette, gently triturate the tissue by pipetting up and down 10-15 times until the solution becomes cloudy and no large clumps remain.
- Plate Cells: Pass the cell suspension through a 70-μm cell strainer to remove any remaining aggregates. Plate the cells onto the pre-coated culture vessels at the desired density. Place the cultures in a 37°C, 5% CO2 incubator [33].
Adult Dorsal Root Ganglia (DRG) Dissociation
Tissue Dissection:
- Animal Source: Sacrifice a 6-week-old young adult rat.
- Expose DRGs: Make a midline incision along the back and carefully remove the spinal column.
- Isolate DRGs: Using a dissection microscope, open the vertebral canal by cutting through the lateral processes. Identify the DRGs attached to the spinal nerves between the vertebrae. Carefully excise them and place in ice-cold DPBS [33].
Enzymatic Digestion:
Mechanical Trituration and Plating:
- Neutralize Enzymes: Add MEM-10 to stop the digestion. Centrifuge at 2000 rpm for 5 minutes and aspirate the supernatant.
- Triturate: Resuspend the pellet in DRG-specific culture medium (F-12 medium supplemented with 10% FBS, Penicillin/Streptomycin, and 20 ng/mL Nerve Growth Factor). Triturate vigorously with a fire-polished Pasteur pipette until a single-cell suspension is achieved.
- Plate Cells: Filter, count, and plate the cells on pre-coated vessels. Maintain cultures in a 37°C, 5% CO2 incubator, refreshing half of the medium every 48 hours [33] [34].
The following workflow diagram visualizes the key decision points and steps in the enzymatic dissociation process for different neural tissues.
Emerging Technologies and Methodological Comparisons
Advanced Non-Enzymatic Dissociation Techniques
Recent technological innovations aim to overcome the limitations of traditional enzymatic methods by employing physical forces for tissue dissociation, thereby preserving cell surface integrity and improving viability for sensitive applications.
Hypersonic Levitation and Spinning (HLS): This contact-free method utilizes a triple-acoustic resonator probe to generate GHz-frequency acoustic waves that create microscale "liquid jets" within the fluid [18]. These jets cause the tissue sample to levitate and undergo a rapid 'press-and-rotate' motion, applying precise hydrodynamic shear forces that disrupt cell-cell connections without direct physical contact. The HLS method reports a 92.3% cell viability and 90% tissue utilization within just 15 minutes for human renal cancer tissue, significantly outperforming traditional methods in speed, yield, and preservation of rare cell populations [18].
Electrical Dissociation: This technique uses applied electric fields to dissociate tissue. One study demonstrated that this method could achieve dissociation of bovine liver tissue and glioblastoma samples in only 5 minutes, yielding over 5 times more cells than traditional enzymatic-mechanical methods [12]. Viability was maintained at approximately 80% for challenging human glioblastoma samples [12].
Cryogenic Enzymatic Dissociation (CED): Developed for challenging samples like Formalin-Fixed Paraffin-Embedded (FFPE) tissues, the CED strategy performs enzymatic digestion with proteinase K at low temperatures [35]. This approach protects the nuclear membrane and retains intranuclear RNA, resulting in a tenfold increase in nuclei yield compared to conventional kits and enhancing gene detection sensitivity in single-nucleus RNA sequencing [35].
Comparative Analysis: Enzymatic vs. Non-Enzymatic Dissociation
The choice between enzymatic and non-enzymatic methods involves trade-offs between yield, viability, surface marker integrity, and technical requirements.
Cellular Viability and Integrity: While optimized enzymatic protocols can achieve high viability, enzymes like trypsin can cleave cell surface receptors and adhesion proteins, potentially altering cell function and signaling responses [8]. Non-enzymatic methods, including HLS and electrical dissociation, generally cause less damage to surface epitopes, better preserving native cellular states for functional assays [18] [8].
Yield and Efficiency: Enzymatic methods typically offer high cell yields and are well-suited for processing large tissue samples. However, advanced non-enzymatic methods are closing this gap. For instance, the HLS system achieves a 90% tissue utilization rate, and electrical dissociation can significantly outperform traditional methods in yield for certain tissues [12] [18].
Technical Considerations and Accessibility: Enzymatic digestion is a cornerstone laboratory technique requiring minimal specialized equipment, making it highly accessible. In contrast, many advanced non-enzymatic methods rely on sophisticated and costly instrumentation (e.g., acoustic resonators, specialized microfluidic devices) and may require extensive protocol optimization for different tissues, posing a barrier to widespread adoption [12] [18] [8].
The diagram below summarizes the core mechanisms and trade-offs of the primary dissociation strategies.
The dissociation of rat cortex, hippocampus, and DRG neurons remains a critical and technically demanding process at the heart of neuroscience research. Evidence-based protocol optimization, as detailed in this guide, is paramount for achieving high neuronal viability, yield, and culture purity. The ongoing methodological evolution from purely enzymatic protocols toward advanced non-enzymatic techniques reflects the field's growing demand for higher fidelity cellular preparations, particularly for single-cell analyses, regenerative medicine, and functional studies.
The future of tissue dissociation lies in the development of standardized, robust, and validated systems that can reproducibly generate high-quality single-cell suspensions from diverse neural tissues with minimal artifacts [12]. While enzymatic methods will likely remain a standard workhorse for routine culture due to their accessibility and efficacy, innovative approaches like Hypersonic Levitation Spinning (HLS) and electrical dissociation show immense promise for applications where preserving native cell surface marker integrity and maximizing viability are paramount [18] [12]. As the cell dissociation market continues to grow, driven by advancements in drug screening and regenerative medicine, we can anticipate increased accessibility and refinement of these advanced technologies, ultimately empowering researchers to obtain deeper and more accurate insights into neuronal function and dysfunction [36].
In neuronal research, the process of detaching adherent cells from culture surfaces is a fundamental yet critical step that can significantly impact cell viability, functionality, and experimental outcomes. Traditional methods have largely relied on enzymatic approaches using trypsin or other proteases, which often damage delicate surface proteins and compromise cellular integrity. Within this context, emerging non-enzymatic technologies offer promising alternatives that preserve the complex architecture and molecular machinery of neuronal cells. This guide provides a comprehensive comparison of three innovative detachment platforms—electrochemical, thermo-responsive, and light-induced systems—evaluating their performance, applications, and implementation requirements for neuroscience research and drug development.
The following table summarizes the key characteristics and performance metrics of the three emerging detachment technologies, based on current research findings.
Table 1: Comparative Performance of Emerging Cell Detachment Technologies
| Technology | Mechanism of Action | Detachment Efficiency | Cell Viability | Detachment Time | Key Applications |
|---|---|---|---|---|---|
| Electrochemical | Alternating current on conductive polymer nanocomposite disrupts adhesion [9] | 95% (osteosarcoma & ovarian cancer cells) [9] | >90% [9] | Minutes [9] | Large-scale biomanufacturing, CAR-T therapies, automated cell culture systems [9] |
| Thermo-responsive | pNIPAAm polymer transition from hydrophobic to hydrophilic below LCST (~32°C) [37] | High (varies by fabrication method) [37] | Preserves cell-cell junctions & ECM [37] | 30 min - 2 hours (varies by method) [37] | Cardiac repair, ocular surface reconstruction, tissue engineering [37] |
| Light-Induced (MXenes) | Photothermal effect converts light to heat, inducing thermal detachment [38] | Research stage (exact efficiency not quantified for cell detachment) [38] | Research stage (not yet quantified for cells) [38] | Ultrafast (theoretical potential) [38] | Potential for optoelectronic devices, thermoelectric harvesting [38] |
Detailed Experimental Protocols
Electrochemical Detachment Method
The electrochemical platform represents one of the most recent advances in non-enzymatic cell detachment, developed by MIT researchers [9]. The methodology employs alternating electrochemical redox-cycling on a nanocomposite biointerface for high-efficiency enzyme-free cell detachment.
Experimental Protocol:
- Surface Preparation: Culture cells on a conductive biocompatible polymer nanocomposite surface engineered specifically for electrochemical responsiveness [9].
- Culture Conditions: Maintain cells under standard conditions until confluence is achieved. For the MIT study, human cancer cells (osteosarcoma and ovarian cancer) were utilized [9].
- Electrical Stimulation: Apply low-frequency alternating voltage to the culture surface. The specific frequency must be optimized for different cell types [9].
- Detachment Monitoring: Observe cell detachment within minutes of stimulation. The MIT team reported detachment efficiency increased from 1% to 95% after identifying optimal frequency parameters [9].
- Cell Collection: Gently collect detached cells while maintaining over 90% viability, as demonstrated in the published results [9].
Thermo-Responsive Detachment Method
Thermo-responsive cell culture surfaces, particularly those grafted with poly(N-isopropylacrylamide) (pNIPAAm), represent the most established non-enzymatic approach, pioneered by Okano's group [37].
Experimental Protocol:
- Surface Fabrication: Prepare thermo-responsive culture surfaces using one of several established methods:
- Electron Beam Polymerization: NIPAAm monomer solution uniformly coated on TCPS and irradiated with 0.3 MGy electron beam [37].
- Plasma Polymerization: Vapor phase plasma polymerization of NIPAAm monomer onto solid surfaces [37].
- UV Irradiation: UV-induced polymerization of NIPAAm monomers on activated surfaces [37].
- Critical Parameters: Ensure pNIPAAm graft thickness is between 15-20 nm for optimal cell attachment and detachment [37].
- Cell Culture: Seed cells on the thermo-responsive surface at 37°C (above LCST), where pNIPAAm is hydrophobic and permits cell adhesion [37].
- Detachment Trigger: Reduce temperature below the LCST (typically to 20°C) for 30 minutes to 2 hours, depending on the specific surface fabrication method and cell type [37].
- Sheet Harvesting: Collect intact cell sheets with preserved extracellular matrix and cell-cell junctions, as demonstrated for various cell types including myocardial cells and epithelial cells [37].
Light-Induced Detachment Using MXenes
While still in earlier stages of development for direct biological applications, MXene materials exhibit properties suitable for light-induced cell detachment through photothermal effects [38].
Experimental Protocol:
- Material Preparation: Synthesize or acquire MXene materials, particularly Ti₃C₂, which demonstrates high efficiency in light-to-heat conversion [38].
- Surface Functionalization: Incorporate MXenes into culture surfaces or utilize them as interfacial layers. Surface terminations significantly influence electronic and thermal properties [38].
- Cell Culture: Allow cells to adhere to the modified surfaces under standard culture conditions.
- Light Activation: Apply specific light wavelengths (based on MXene optical properties) to trigger photothermal heating. Ultrafast spectroscopic techniques have revealed the fundamental charge and thermal transport effects in these materials [38].
- Thermal Detachment: Utilize the generated heat to disrupt cell-surface interactions. The unusual combination of high electrical mobility and intrinsically low thermal conductivity in MXenes offers exciting opportunities for controlled thermal applications [38].
Technology Workflows
The following diagram illustrates the operational workflow for each detachment technology, highlighting key decision points and procedural steps:
Diagram 1: Operational workflows for the three detachment technologies
Research Reagent Solutions
Implementation of these emerging technologies requires specific materials and reagents, as detailed below:
Table 2: Essential Research Reagents for Emerging Detachment Technologies
| Technology | Key Materials | Function/Purpose | Implementation Considerations |
|---|---|---|---|
| Electrochemical | Conductive polymer nanocomposite [9] | Provides electroactive surface for cell culture and detachment | Requires specialized fabrication; compatible with various cell types |
| Thermo-responsive | pNIPAAm (poly(N-isopropylacrylamide)) [37] | Temperature-responsive polymer that changes wettability | Graft thickness critical (15-20 nm optimal); multiple fabrication methods available |
| Thermo-responsive | NIPAAm monomer [37] | Precursor for pNIPAAm surface grafting | purity essential for consistent performance |
| Thermo-responsive | Tissue culture polystyrene (TCPS) [37] | Standard substrate for pNIPAAm grafting | Compatible with electron beam and plasma polymerization |
| Light-Induced | MXene (Ti₃C₂) materials [38] | Photothermal conversion material | Surface terminations dictate properties; requires controlled synthesis |
| General | Cell culture media without Ca²⁺/Mg²⁺ | Facilitates detachment by reducing integrin-mediated adhesion | Common to multiple non-enzymatic approaches |
Discussion & Research Implications
Advantages and Limitations
Each technology presents distinct advantages for neuronal research applications. The electrochemical approach offers rapid detachment with high viability, making it suitable for time-sensitive applications and automated systems [9]. Its compatibility with large-scale biomanufacturing addresses a critical need in therapeutic development. However, the requirement for specialized conductive surfaces may increase implementation costs.
The thermo-responsive method provides the unique advantage of harvesting intact cell sheets with preserved extracellular matrix and cell-cell junctions [37]. This is particularly valuable for tissue engineering and transplantation applications where structural integrity is essential. The main limitations include relatively slow detachment times (30 minutes to 2 hours) and the need for precise temperature control systems.
The light-induced approach using MXenes offers theoretical potential for ultrafast, spatially controlled detachment [38]. The photothermal properties of MXenes enable precise manipulation, but this technology remains in earlier stages of development for biological applications. Further research is needed to establish protocols and validate biocompatibility.
Considerations for Neuronal Research
For neuronal researchers, selection of detachment methods requires careful consideration of experimental goals. Primary neurons are particularly sensitive to surface protein damage, making non-enzymatic approaches preferable. Studies have shown that enzymatic methods can compromise surface proteins like Fas ligands and Fas receptors, requiring up to 20 hours for recovery [17]. This protein damage can significantly impact neuronal signaling studies and receptor function investigations.
When transitioning from traditional methods, researchers should note that enzyme-free dissociation buffers have demonstrated lower cell viability (68.7%) compared to trypsin (93.2%) in mesenchymal stem cells [16]. However, the emerging technologies discussed herein aim to overcome these limitations through innovative mechanisms that preserve cellular integrity while enabling efficient detachment.
The evolving landscape of cell detachment technologies offers neuroscience researchers powerful tools that overcome the limitations of traditional enzymatic methods. Electrochemical systems provide rapid, high-viability detachment suitable for automated workflows; thermo-responsive approaches enable preservation of complex cellular architectures; while light-induced methods present opportunities for precise spatiotemporal control. Selection among these platforms should be guided by specific research requirements, considering factors of scale, speed, structural preservation, and implementation practicality. As these technologies continue to mature, they hold significant promise for advancing neuronal research, drug development, and regenerative medicine applications.
Solving Common Challenges in Neuronal Detachment and Culture
In neuroscience research, the process of detaching adherent cells for subculturing or analysis is a fundamental but critical step. The choice between enzymatic and non-enzymatic detachment methods significantly influences experimental outcomes by directly impacting the integrity of surface receptors and functional proteins. Proteolytic damage during cell harvesting can cleave vital neuronal surface markers, receptors, and adhesion proteins, thereby altering cell signaling, viability, and functionality. This guide provides an objective comparison of these methodologies, focusing on their effects on cell health, surface protein preservation, and downstream applications in neuronal research. Understanding these trade-offs is essential for researchers aiming to maintain physiological relevance in their experimental models while achieving efficient cell dissociation.
Comparative Analysis of Detachment Methodologies
The fundamental difference between enzymatic and non-enzymatic methods lies in their mechanism of action. Enzymatic methods use proteases like trypsin or Accutase to actively cleave peptide bonds in cell-adhesion proteins and the extracellular matrix. In contrast, non-enzymatic methods typically rely on chelating agents like EDTA that bind calcium and magnesium ions, disrupting calcium-dependent cell adhesions without proteolytic cleavage [16] [17] [8].
The tables below summarize core characteristics and experimental outcomes for both approaches.
Table 1: Fundamental Characteristics of Detachment Methods
| Feature | Enzymatic Methods | Non-Enzymatic Methods |
|---|---|---|
| Mechanism | Proteolytic cleavage of adhesion proteins and ECM [8] | Chelation of Ca²⁺/Mg²⁺ ions, disrupting cell-cell and cell-ECM interactions [16] [17] |
| Primary Agents | Trypsin, TrypLE, Accutase, collagenase | EDTA-based buffers, PBS-based dissociation buffers |
| Key Advantage | Rapid and efficient detachment, even for strongly adherent cells [8] | Preserves structural integrity of membrane surface proteins [16] |
| Key Limitation | Degrades surface proteins and glycoproteins; can alter cell metabolism and viability [16] [39] | Slower dissociation; may be insufficient for strongly adherent cells; can lower reattachment efficiency [16] |
Table 2: Comparative Experimental Outcomes on Cell Health and Function
| Experimental Metric | Enzymatic Methods | Non-Enzymatic Methods | Key Research Findings |
|---|---|---|---|
| Cell Viability Post-Detachment | Higher viability reported (e.g., 93.2% with trypsin) [16] | Lower viability reported (e.g., 68.7% with enzyme-free buffer) [16] | Trypsin yielded a significantly higher proportion of viable MSCs [16] |
| Reattachment Efficiency | Superior reattachment rates post-dissociation [16] | Significantly lower proportion of viable cells reattach [16] | Critical for experiments requiring continued culture after passaging [16] |
| Surface Protein Integrity | Can compromise specific surface proteins (e.g., FasL, Fas receptor) [17] | Better preservation of many surface protein epitopes [16] | Accutase cleaves FasL and Fas receptor, requiring a 20-hour recovery period [17] |
| Detachment Time | Relatively fast (e.g., ~5-6 minutes for MSC monolayers) [16] | Generally slower (e.g., ~15-16 minutes for MSC monolayers) [16] | Efficiency can vary significantly by cell type and confluency [16] |
Detailed Experimental Protocols and Data
To ensure reproducibility, this section outlines key methodologies from cited studies and presents quantitative results.
Protocol: Comparing Trypsin and Enzyme-Free Dissociation Buffer
This protocol is adapted from a study comparing the dissociation of adherent mesenchymal stem cell (MSC) monolayers [16].
- Step 1: Cell Preparation: Culture confluent monolayers of MSC in 12-well cell culture dishes.
- Step 2: Pre-Treatment: Wash monolayers two times with Ca²⁺-free PBS prior to dissociation.
- Step 3: Detachment: Add 1 ml of pre-warmed (37°C) 0.05% (w/v) Trypsin-EDTA or enzyme-free, PBS-based cell dissociation buffer to each well.
- Step 4: Incubation: Place dishes in a cell culture incubator. Gently pipet the solution every 2-3 minutes.
- Step 5: Termination & Collection: Once cells detach, collect the cell suspension into microcentrifuge tubes and centrifuge at 500×g for 5 minutes.
- Step 6: Analysis: Discard supernatant, reconstitute cell pellet in 0.5 ml PBS (with Ca²⁺), and analyze cell viability using trypan blue exclusion assay with an automated cell counter.
- Step 7: Reattachment Assay: Re-seed dissociated cells into new culture dishes. After 24 hours, wash off unattached cells and perform an MTT assay on the reattached cells to quantify metabolic activity [16].
Protocol: Assessing Surface Protein Integrity Post-Detachment
This protocol is adapted from a study investigating the effect of accutase on surface proteins in macrophages [17].
- Step 1: Cell Detachment: Treat adherent cells (e.g., RAW264.7 macrophages) with either an EDTA-based non-enzymatic detachment solution or accutase for 10-30 minutes, as per manufacturer's instructions.
- Step 2: Recovery Culture (Optional): For recovery analysis, reseed a portion of the detached cells in complete medium and culture for 2 to 20 hours.
- Step 3: Flow Cytometry Analysis: Harvest cells at designated time points (immediately after detachment or after recovery). Stain cells with fluorescently-labeled antibodies against target surface proteins (e.g., FasL, Fas receptor) and appropriate isotype controls.
- Step 4: Data Acquisition and Analysis: Analyze stained cells using a flow cytometer. Compare the Mean Fluorescence Intensity (MFI) of stained cells between detachment groups to quantify surface protein levels [17].
Quantitative Results from Key Studies
The following table synthesizes core quantitative findings from the referenced research, providing a direct comparison of outcomes.
Table 3: Synthesis of Key Experimental Data
| Study & Metric | Enzymatic Method Result | Non-Enzymatic Method Result | Notes |
|---|---|---|---|
| MSC Viability [16] | 93.2% ± 3.2% (Trypsin) | 68.7% ± 5.0% (Enzyme-free buffer) | p = 0.002 |
| MSC Reattachment [16] | Significantly higher | Significantly lower | Measured by MTT assay 24h post-seeding |
| Surface FasL MFI [17] | Significantly decreased (Accutase) | Preserved (EDTA-based solution) | Effect was reversible after 20h recovery |
| Detachment Time [16] | ~5-6 minutes (Trypsin) | ~15-16 minutes (Enzyme-free buffer) | For confluent MSC monolayers |
Visualization of Mechanisms and Workflows
Detachment Method Impact on Cell Surface
This diagram illustrates the fundamental mechanisms of how enzymatic and non-enzymatic detachment methods affect surface proteins and subsequent cell recovery.
Experimental Workflow for Method Comparison
This diagram outlines a generalized experimental workflow for comparing the effects of different detachment methods on cells, suitable for adaptation in neuronal research.
The Scientist's Toolkit: Essential Research Reagents
Selecting appropriate reagents is fundamental to designing robust experiments. The following table details key solutions used in the studies cited in this guide.
Table 4: Key Reagent Solutions for Cell Detachment Studies
| Reagent / Solution | Function / Description | Example Application Context |
|---|---|---|
| Trypsin-EDTA | Proteolytic enzyme that cleaves peptide bonds; EDTA chelates ions to enhance detachment efficiency [16] [8]. | General cell culture passaging where surface protein integrity is not the primary concern [16]. |
| Enzyme-Free Dissociation Buffer | Isotonic, PBS-based solution containing salts and chelating agents to disrupt cell adhesion without proteolysis [16]. | Experiments requiring preservation of surface epitopes for flow cytometry or immunohistochemistry [16]. |
| Accutase | A mixture of proteolytic and collagenolytic enzymes, considered milder than trypsin for many cell types [17]. | Detachment of sensitive cells; however, requires validation for specific surface proteins of interest [17]. |
| MTT Reagent | (3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide); used to assess metabolic activity of cells as a proxy for viability and proliferation [16]. | Quantifying reattachment efficiency and metabolic health of cells after detachment and re-seeding [16]. |
| Trypan Blue Solution | A vital dye used to stain non-viable cells, allowing for quantification of cell viability post-detachment [16]. | Standard, rapid assessment of immediate cell death caused by the detachment process [16]. |
The choice between enzymatic and non-enzymatic cell detachment methods presents a clear trade-off. Enzymatic methods, particularly trypsin, offer speed and efficiency, resulting in higher initial cell viability and superior reattachment rates for some cell types like MSCs [16]. However, this comes at the cost of potential proteolytic damage to surface receptors, which can profoundly impact neuronal signaling studies and requires significant recovery time [17] [39]. Non-enzymatic methods excel at preserving surface protein integrity, crucial for immunophenotyping and functional studies, but may be less effective for strongly adherent cells and can result in lower viability and reattachment efficiency [16] [8].
For neuronal research, the optimal method depends heavily on the specific experimental endpoint. Studies prioritizing immediate post-detachment analysis of surface markers benefit from non-enzymatic approaches. In contrast, experiments requiring large numbers of healthy, proliferating cells after passaging might favor enzymatic methods, provided adequate time is allowed for surface protein recovery. Researchers must weigh these factors carefully, as the detachment process is not merely a technical step but a critical variable that can define the physiological relevance and success of subsequent investigations.
The process of detaching neurons from culture surfaces is a fundamental yet critical step in neuroscience research, cell therapy manufacturing, and regenerative medicine. The choice between enzymatic and non-enzymatic detachment methods carries significant implications for cell viability, surface marker integrity, and subsequent experimental outcomes. Enzymatic treatments, particularly trypsin, can damage delicate cell membranes and surface proteins, while non-enzymatic approaches may reduce viability and require longer processing times. Understanding the recovery timelines for neurons to regenerate these surface markers post-detachment is essential for obtaining reliable data, particularly for flow cytometry analyses and functional studies that depend on surface antigen expression. This guide provides a comprehensive comparison of detachment methods and their impact on neuronal surface markers, synthesizing current evidence to inform research protocols and experimental design.
Detachment Method Comparison: Mechanisms and Impacts
Enzymatic vs. Non-Enzymatic Dissociation
The fundamental differences between dissociation methods directly impact neuronal surface marker integrity and subsequent recovery needs.
Table 1: Comparison of Cell Detachment Methods
| Method Type | Specific Agents | Mechanism of Action | Processing Time | Key Advantages | Key Limitations |
|---|---|---|---|---|---|
| Enzymatic | Trypsin | Proteolytic digestion of adhesion proteins | 5-6 minutes [16] | High cell viability (93.2%) [16] | Damages surface proteins [23] |
| Enzymatic | TrypLE | Recombinant microbial enzyme | ~11-17 minutes [21] | Reduced animal-derived components | Variable cleavage of surface markers [23] |
| Enzymatic | Accutase | Enzyme mixture | Varies | Gentler than trypsin | Can cleave M2 markers CD206, CD163 [23] |
| Non-Enzymatic | EDTA | Chelates Ca²⁺/Mg²⁺ ions | 15-16 minutes [16] | Preserves surface protein integrity [16] | Lower viability (68.7%) [16] |
| Non-Enzymatic | Cell Scraping | Mechanical force | Immediate | Rapid, cost-effective | Irreversible cell damage [9] |
| Electrochemical | Alternating current | Disrupts adhesion via redox cycling | Minutes [9] | High viability (>90%), enzyme-free [9] | Requires specialized surfaces [9] |
The primary challenge with enzymatic methods is their proteolytic action on surface markers essential for neuronal identification and sorting. Studies demonstrate that enzymatic detachment not only cleaves surface proteins but does so selectively, with variable effects across different markers and donors [23]. For instance, Accutase has been shown to selectively cleave the M2 macrophage markers CD206 and CD163, complicating the study of these specific cell populations [23].
Non-enzymatic alternatives address the surface marker damage but introduce different limitations. EDTA and other chelating agents work by sequestering divalent cations necessary for cell adhesion, but result in significantly lower cell viability (68.7% vs 93.2% for trypsin) and reduced reattachment capacity [16]. Mechanical methods like scraping cause direct physical damage to cells and are not suitable for closed systems or multi-layered vessels [9].
Emerging technologies like electrochemical detachment offer promising alternatives by using low-frequency alternating current on conductive biocompatible polymer surfaces to disrupt cell adhesion without enzymatic action, achieving over 90% viability and high detachment efficiency [9]. This method could potentially minimize surface marker damage while maintaining high cell viability.
Experimental Protocols for Neuronal Detachment and Analysis
Standardized Detachment Workflow
The following protocol outlines a standardized approach for detaching neuronal cultures while preserving surface marker integrity for subsequent analysis:
1. Pre-harvest Assessment
- Examine cultures using phase-contrast microscopy to assess confluence and overall cell health prior to detachment [40].
- Document baseline morphology as a reference for post-detachment recovery assessment.
2. Harvesting Procedure
- Gently wash adherent neural cells with Mg²⁺/Ca²⁺-free phosphate-buffered saline (PBS) at room temperature [40].
- Add pre-warmed (37°C) detachment solution (choice dependent on experimental priorities).
- Incubate at 37°C for optimized duration:
- Gently tap vessel or pipette to facilitate cell detachment, avoiding excessive mechanical stress [40].
3. Reaction Quenching and Cell Collection
- Quench enzymatic activity using 2x volume of flow buffer (PBS with 2% FBS) [40].
- Collect cell suspension in 15mL conical tubes.
- Centrifuge at 500×g for 5 minutes [16].
- Discard supernatant and resuspend pellet in appropriate culture medium or buffer.
4. Post-Detachment Processing
- Assess viability using trypan blue exclusion assay [16].
- Count cells using automated cell counter or hemocytometer.
- Proceed with surface marker analysis or return to culture for recovery studies.
Surface Marker Analysis via Flow Cytometry
For analyzing neuronal surface markers post-detachment, the following flow cytometry protocol is recommended:
Surface Antigen Staining
- Resuspend harvested cells in flow buffer (PBS with 2% FBS) at appropriate concentration.
- Aliquot cells into staining tubes (approximately 1×10⁶ cells per tube).
- Add conjugated CD antibodies (e.g., CD24, CD44, CD184) at determined optimal concentrations.
- Incubate for 30 minutes at 4°C in the dark.
- Wash cells twice with flow buffer by centrifugation at 500×g for 5 minutes.
- Resuspend in flow buffer for immediate analysis or fix with 1-4% paraformaldehyde if analysis will be delayed [40].
Intracellular Antigen Staining (if required)
- After surface staining, fix cells with 4% PFA for 15 minutes at room temperature.
- Permeabilize cells with 0.1-0.5% Triton X-100 in PBS for 10 minutes.
- Incubate with primary antibody against intracellular target for 1 hour at room temperature.
- Wash twice with permeabilization buffer.
- Incubate with fluorescent-conjugated secondary antibody for 30 minutes at room temperature.
- Wash twice with flow buffer and resuspend for analysis [40].
Flow Cytometry Analysis
- Use appropriate gating strategies to identify target populations:
- Forward scatter (FSC) vs side scatter (SSC) to exclude debris
- Single cells using FSC-A vs FSC-H
- Viable cells using viability dye exclusion
- Fluorescence minus one (FMO) controls for setting positive populations [40]
Neuronal Surface Marker Signatures and Detachment Impact
Key Neuronal Surface Markers
Different neural cell types express distinct surface marker signatures that can be used for identification and purification:
Table 2: Neural Cell Surface Marker Signatures
| Cell Type | Surface Marker Signature | Function/Importance | Impacted by Detachment Methods |
|---|---|---|---|
| Neural Stem Cells (NSC) | CD184+/CD271−/CD44−/CD24+ [41] | Identifies multipotent stem cells capable of generating neurons and glia [41] | Enzymatic methods may cleave CD24 and CD184, affecting sorting efficiency |
| Neurons | CD184−/CD44−/CD15LOW/CD24+ [41] | Identifies post-mitotic neurons; CD24 is a cell adhesion molecule [41] | CD24 particularly vulnerable to enzymatic cleavage |
| Glia | CD184+/CD44+ [41] | Identifies glial populations including astrocytes [41] | CD44 may be affected by prolonged enzymatic treatment |
| Neural Stem Cells | CD133+/CD15+ [41] | Alternative NSC signature; CD133 is a glycoprotein stem cell marker [41] | Glycoprotein markers sensitive to enzymatic degradation |
The recovery of these surface markers post-detachment is crucial for accurate identification and sorting of neural populations. Current evidence suggests that the regeneration timeline for surface markers depends on multiple factors:
- Extent of initial cleavage: More extensive proteolysis requires longer recovery
- Marker turnover rate: Inherent synthesis and membrane integration rates
- Cellular health post-detachment: Viable cells with proper metabolic function regenerate markers more efficiently
- Culture conditions: Growth factors and extracellular matrix support influence recovery
While specific quantitative timelines for neuronal surface marker regeneration are not well-documented in the literature, general observations indicate that:
- Mild enzymatic treatment may require 24-48 hours for full surface marker regeneration
- After extensive proteolysis, complete recovery may take 72 hours or longer
- Non-enzymatic methods typically require less recovery time as surface integrity is preserved
Research Reagent Solutions Toolkit
Table 3: Essential Reagents for Neuronal Detachment and Analysis
| Reagent Category | Specific Products | Primary Function | Considerations for Neuronal Research |
|---|---|---|---|
| Enzymatic Detachment | Trypsin-EDTA [16] | Proteolytic dissociation | Rapid but damages surface markers; use at 0.05% concentration [16] |
| Enzymatic Detachment | TrypLE Express [21] | Recombinant enzyme alternative | Reduced animal-derived components; gentler than trypsin [21] |
| Enzymatic Detachment | Accutase [23] | Enzyme mixture | Gentler on surface markers but still cleaves specific targets [23] |
| Non-Enzymatic Detachment | EDTA-based buffer [16] | Chelation of divalent cations | Preserves surface markers but reduces viability [16] |
| Flow Cytometry | CD24, CD44, CD184 antibodies [41] | Neural population identification | Essential for sorting NSC, neurons, and glia [41] |
| Viability Assessment | Trypan blue [16] | Cell viability staining | Use with automated cell counter for consistency [16] |
| Cell Culture | MSCGM bullet kit [16] | Culture medium for mesenchymal cells | Optimized for stem cell growth and recovery |
| Analysis | MTT reagent [16] | Metabolic activity assay | Assess recovery post-detachment through metabolic function [16] |
Method Selection Guidelines
Strategic Decision Framework
Choosing the appropriate detachment method requires balancing multiple factors specific to research goals:
Application-Specific Recommendations:
- Flow cytometry and cell sorting: Prioritize non-enzymatic methods or brief enzymatic exposure followed by 24-48 hour recovery period
- Electrophysiological studies: Enzymatic methods with controlled duration to preserve functional properties
- Cell transplantation: Balance viability and surface marker integrity using optimized enzymatic protocols or electrochemical methods
- RNA sequencing: Minimize enzymatic exposure to prevent artifactual gene expression changes
- Long-term culture: Use gentlest effective method to preserve proliferative capacity and differentiation potential
The recovery of neuronal surface markers post-detachment represents a critical yet understudied aspect of neural cell culture. Current evidence indicates that enzymatic methods, particularly trypsin, efficiently dissociate cells but significantly damage surface markers, potentially requiring 24-72 hours for complete regeneration. Non-enzymatic methods preserve surface marker integrity but compromise viability and yield. Emerging technologies like electrochemical detachment offer promising alternatives with high viability and minimal surface protein damage.
Researchers should select detachment methods based on their specific endpoint applications, build in appropriate recovery periods when surface marker integrity is crucial, and consistently report detachment protocols and recovery times in publications to advance our understanding of neuronal surface marker regeneration. As the field moves toward more standardized, automated, and gentle dissociation technologies, the variability introduced by current detachment methods should decrease, enhancing reproducibility across neural research applications.
Preventing Low Viability and Poor Re-attachment in Sensitive Primary Cultures
The success of in vitro neuroscience research heavily relies on the ability to isolate and culture primary neurons that maintain high viability and robust re-attachment capabilities. The initial step of detaching cells from tissue or culture surfaces is particularly critical, as the chosen method can directly dictate experimental outcomes. This guide provides a comparative analysis of enzymatic versus non-enzymatic detachment methods, framing them within the broader context of optimizing workflows for sensitive primary neuronal cultures. We objectively evaluate performance based on cell viability, re-attachment efficiency, and functional preservation, providing the data and protocols necessary for researchers to make informed decisions that enhance the reproducibility and reliability of their findings.
Method Comparison: Enzymatic vs. Non-Enzymatic Dissociation
The choice between enzymatic and non-enzymatic dissociation is a fundamental decision point. The table below summarizes a performance comparison based on aggregated experimental data.
Table 1: Performance Comparison of Cell Detachment Methods for Neuronal Cultures
| Method | Typical Viability | Typical Re-attachment Efficiency | Key Advantages | Key Disadvantages | Ideal Use Cases |
|---|---|---|---|---|---|
| Enzymatic (e.g., Trypsin) | ~90-93% [42] | Significantly higher [42] | Rapid; effective for strongly adherent cells; high cell yield [43] [44] | Can damage cell surface proteins [9] [12] | Large-scale expansion; cultures where surface protein integrity is not critical [43] |
| Non-Enzymatic (Chemical) | ~69% [42] | Significantly lower [42] | Gentle on surface proteins; animal-origin free [44] | Slower; less effective for strongly adherent cells; lower yield [44] [42] | Flow cytometry; receptor studies [44] |
| Mechanical | ~33% [43] | Not Specified | Simple; no chemical exposure | Low viability and yield; high shear stress [43] | Not recommended for sensitive primary neurons [43] |
| Emerging (Electrochemical) | >90% [9] | Not Specified (Promising for automation) | Enzyme-free; high viability; suitable for automation [9] | Early-stage technology | Potential for GMP-compliant cell therapy manufacturing [9] |
Detailed Experimental Data and Protocols
To ensure reproducibility, this section outlines specific experimental workflows and the resulting quantitative data for the primary methods discussed.
Experimental Protocol: Enzymatic Dissociation of Primary Embryonic Spinal Cord Neurons
The following protocol, adapted from Jiang et al. (2006), is an example of an optimized enzymatic process for high-yield isolation of primary neurons [43].
- Tissue Source: Spinal cords from E14-15 rat embryos.
- Dissociation Solution: Enzymatic solution (e.g., trypsin or TrypLE [44]).
- Procedure:
- Dissect spinal cords and remove meninges carefully.
- Mince tissue into 3-4 mm pieces.
- Wash tissue pieces in a balanced salt solution without calcium and magnesium.
- Incubate tissue with the pre-warmed enzymatic solution at 37°C for a defined period (e.g., 20-30 minutes for trypsin [44]), with gentle agitation.
- Terminate the enzymatic reaction by adding complete medium containing serum or a trypsin inhibitor.
- Triturate the tissue gently but thoroughly with a pipette to create a single-cell suspension.
- Filter the suspension through a sterile mesh (70-100 µm) to remove clumps.
- Centrifuge the cell suspension and resuspend the pellet in complete neuronal culture medium.
- Count cells and assess viability using trypan blue exclusion before plating.
- Key Parameters for Success:
- Embryonic Age: Critically impacts neuronal yield and survival (E14-15 recommended for spinal cord) [43].
- Seeding Density: Must be optimized for the specific neuronal type and assay [43] [33].
- Coating: Culture surfaces must be pre-coated with substrates like poly-L-ornithine/laminin or Polyethylenimine (PEI) to promote attachment and neurite outgrowth [11].
Experimental Workflow: Cell Detachment from Culture Surfaces
The following diagram illustrates the core decision-making workflow for selecting a detachment method when subculturing adherent neuronal cells.
Key Supporting Data
The comparative study by Jiang et al. provides direct, quantitative evidence for the superiority of enzymatic over mechanical dissociation for primary tissue [43]:
- Cell Yield: Enzymatic dissociation yielded 25.40 ± 5.41 × 10⁶ cells per 12 embryos, compared to only 3.43 ± 0.52 × 10⁶ cells via mechanical means.
- Cell Viability: Viability was significantly higher with the enzymatic method (83.40 ± 3.08%) versus mechanical dissociation (32.81 ± 3.49%).
For dissociated cells in culture, a study on Mesenchymal Stem Cells (MSCs) highlights a critical trade-off. While trypsin yielded 93.2% viability and significantly higher re-attachment, enzyme-free buffer resulted in 68.7% viability and poor re-attachment, underscoring the method's impact on subsequent culture health [42].
The Scientist's Toolkit: Essential Reagents & Materials
Successful culture of primary neurons depends on a suite of specialized reagents. The following table details essential items and their functions.
Table 2: Essential Research Reagents for Primary Neuronal Culture
| Reagent/Material | Function | Example Use Case |
|---|---|---|
| Trypsin | Proteolytic enzyme for digesting cell-adhesion proteins. Effective for strongly adherent cells [44]. | Standard dissociation of primary tissues and strongly adherent neuronal cell lines [43] [44]. |
| TrypLE Express | Recombinant, animal-origin-free enzyme. Functions as a direct substitute for trypsin [44]. | Ideal for therapeutic cell manufacturing where animal-derived components must be avoided [44]. |
| Cell Dissociation Buffer (Non-Enzymatic) | Chelates calcium and magnesium to disrupt cell-cell and cell-matrix adhesion. Gentle on surface proteins [44] [42]. | Gently detaching lightly adherent cells or when preserving surface receptor integrity is paramount [44]. |
| Poly-L-Ornithine/Laminin | Synthetic peptide/natural protein coating for culture surfaces. Promotes neuronal attachment and neurite outgrowth [11]. | Standard coating combination for primary neurons from cortex, hippocampus, and spinal cord [33] [11]. |
| Polyethylenimine (PEI) | Synthetic polymer coating resistant to proteolysis. Promotes even cell distribution and strong attachment [11]. | Coating for MEA plates to reduce cell aggregation and improve signal detection in electrophysiology studies [11]. |
| Neurobasal Medium with B-27 | Serum-free medium optimized for long-term survival of primary neurons. Suppresses glial cell growth [33]. | Primary culture of central nervous system neurons (e.g., cortical, hippocampal) [33]. |
Mechanisms of Action and Impact on Cells
Understanding how detachment methods work at a cellular level helps predict their impact on experimental outcomes. The following diagram contrasts the mechanisms of enzymatic and non-enzymatic methods.
Selecting the optimal cell detachment method is a critical step that cannot be reduced to a one-size-fits-all approach. For researchers requiring maximum cell yield and viability from primary tissue or robust subculturing, enzymatic methods like trypsin or TrypLE currently present the most effective and reliable option, despite their potential to cleave surface proteins [43] [42]. Conversely, when the experimental endpoint demands intact surface markers, non-enzymatic chelation buffers are the necessary choice, albeit with a trade-off in viability and re-attachment efficiency [42]. Emerging technologies, such as electrochemical detachment, offer a promising future alternative by potentially decoupling high viability from enzymatic damage [9]. The key to preventing low viability and poor re-attachment lies in aligning the detachment strategy with the specific cellular material and ultimate goals of the research.
Cell detachment is a fundamental step in neuronal research, enabling cell propagation, analysis, and application in therapeutic development. The choice between enzymatic and non-enzymatic detachment methods significantly impacts experimental outcomes by influencing cell viability, surface marker integrity, functional properties, and downstream application success. This guide provides a systematic comparison of detachment strategies, supported by experimental data, to help researchers select the optimal method for specific neuronal applications. The growing cell dissociation market, projected to reach USD 1621.47 million by 2035 at a CAGR of 13.5%, reflects increasing emphasis on standardized, reliable cell processing methods across biopharmaceutical and research sectors [45].
Understanding the mechanisms of cell adhesion and detachment provides the foundation for method selection. Adherent cells, including many neuronal models, attach to surfaces via integrin-mediated binding to extracellular matrix proteins, cadherin-based cell-cell junctions, and other specialized adhesion complexes [8]. Detachment methods work by disrupting these interactions through proteolytic cleavage (enzymatic methods) or modulation of ionic and physical interactions (non-enzymatic methods), each with distinct implications for cell integrity and function.
Methodological Comparison: Experimental Protocols and Performance Metrics
Quantitative Comparison of Detachment Methods
Table 1: Comprehensive Performance Metrics of Cell Detachment Methods
| Method | Cell Viability | Detachment Time | Surface Marker Preservation | Post-Detachment Function | Best Applications |
|---|---|---|---|---|---|
| Trypsin | 93.2% [16] | 5-6 minutes [16] | Low (cleaves surface proteins) [8] [17] | Reduced reattachment capacity [16] | Large-scale expansion, routine passaging |
| Accutase | High (maintains viability >90 minutes) [17] | 10-30 minutes [17] | Variable (cleaves FasL/Fas receptors) [17] | Requires 20h recovery for marker reexpression [17] | Gentle dissociation for sensitive cells |
| Non-enzymatic Buffer | 68.7% [16] | 15-16 minutes [16] | High (preserves surface markers) [17] [46] | Lower reattachment rates [16] | Flow cytometry, immunostaining |
| Scraping | Variable (mechanical damage risk) | Immediate | High (no chemical alteration) [17] | Maintains function but risk of activation [46] | Protein analysis, RNA studies |
| Electrochemical | >90% [9] | Minutes [9] | Expected high (enzyme-free) | High functionality for therapies [9] | CAR-T therapies, sensitive primary cells |
| Thermoresponsive Surfaces | High [47] | 5-10 minutes [47] | High (physical detachment) | Maintains differentiation potential [8] | Stem cell research, tissue engineering |
Detailed Experimental Protocols
Enzymatic Detachment with Trypsin
Protocol:
- Aspirate culture medium and wash cells with Ca2+-free PBS [16].
- Add pre-warmed 0.05% (w/v) Trypsin-EDTA solution [16].
- Incubate at 37°C for 5-6 minutes with gentle pipetting every 2-3 minutes [16].
- Neutralize with complete medium containing serum [8].
- Centrifuge at 500×g for 5 minutes and resuspend in fresh medium [16].
Critical Parameters:
- Concentration: Standard trypsin at 0.05% balances efficiency and cell damage [16].
- Temperature: 37°C incubation optimizes enzyme activity [8].
- Monitoring: Visual inspection for cell rounding and detachment prevents over-exposure [21].
Non-enzymatic Detachment with EDTA-Based Buffer
Protocol:
- Remove culture medium and wash with PBS [16].
- Add enzyme-free dissociation buffer (isotonic, PBS-based with chelating agents) [16].
- Incubate at 37°C for 15-16 minutes with periodic pipetting [16].
- Collect cell suspension and centrifuge at 500×g for 5 minutes [16].
- Resuspend in appropriate medium for downstream applications [46].
Critical Parameters:
- Composition: Calcium and magnesium-free PBS with EDTA concentrations approximately 0.53 mM [16].
- Temperature: 37°C incubation enhances efficiency [16].
- Mechanical assistance: Gentle pipetting necessary due to slower action [16].
Lens-Free Imaging Monitoring Protocol
Novel Monitoring Approach:
- Set up lens-free imaging device inside cell culture incubator (37°C, 5% CO2) [21].
- Calibrate on cells in PBS prior to dissociation reagent application [21].
- Start time-lapse experiment with 20-second intervals during detachment process [21].
- Reconstruct phase and intensity images using iterative phase recovery method [21].
- Automatically extract detachment percentage using image analysis software [21].
- Inhibit enzymatic reaction when approximately 92.5% of cells are detached [21].
Validation: This method achieved average absolute error values of 1.49-1.97% across different cell densities, providing robust quantification of detachment progression [21].
Molecular Mechanisms: Signaling Pathways and Functional Impacts
Detachment-Induced Signaling Pathways
Diagram 1: Molecular signaling pathways activated during cell detachment. Enzymatic methods (left) trigger proteolysis leading to surface protein cleavage and signaling alterations, while non-enzymatic methods (right) preserve surface markers and maintain normal cellular function.
Method Selection Decision Framework
Diagram 2: Decision framework for selecting detachment methods based on research priorities and applications. This workflow guides researchers in matching detachment strategies to specific experimental requirements.
Research Reagent Solutions: Essential Materials for Cell Detachment
Table 2: Key Research Reagents and Solutions for Cell Detachment Applications
| Reagent/Solution | Composition | Mechanism of Action | Primary Applications | Key Considerations |
|---|---|---|---|---|
| Trypsin-EDTA | Proteolytic enzyme (trypsin) + chelating agent (EDTA) in buffer | Proteolytic cleavage of adhesion proteins + calcium chelation | Routine cell culture, large-scale expansion | Concentration-dependent cytotoxicity; requires neutralization [8] [16] |
| Accutase | Blend of proteolytic and collagenolytic enzymes in PBS | Gentle enzymatic degradation of matrix proteins | Sensitive cell types, stem cells, primary cultures | Cleaves specific surface markers (FasL/Fas); requires recovery time [17] |
| Enzyme-Free Dissociation Buffer | Isotonic salt solution with chelating agents | Chelation of Ca2+/Mg2+ ions required for integrin binding | Surface marker analysis, flow cytometry, functional studies | Slower action; may require mechanical assistance [16] |
| Thermoresponsive Polymers | PNIPAM coatings with nanoscale architecture [47] | Temperature-modulated hydration/swelling generates disjoining pressure | Cell sheet engineering, therapeutic applications | Requires specialized surfaces; limited to compatible culture vessels [47] |
| Electrochemical Platforms | Conductive polymer nanocomposite surfaces [9] | Alternating current disrupts cell-adhesion interactions | Automated systems, high-throughput applications, CAR-T manufacturing | Emerging technology; requires specialized equipment [9] |
| Lens-Free Imaging Systems | CMOS sensor, LED illumination, computational reconstruction [21] | Real-time monitoring of detachment via optical feature analysis | Process optimization, standardization, quality control | Enables precise endpoint determination [21] |
Application-Specific Recommendations for Neuronal Research
Primary Neuronal Culture and Functional Studies
For primary neuronal cultures, non-enzymatic methods or gentle enzymatic alternatives like Accutase are recommended. Research demonstrates that enzymatic methods can cause cleavage of surface receptors critical for neuronal function and signaling [17]. When studying neuronal receptor distribution, signaling pathways, or synaptic function, EDTA-based detachment or specialized non-enzymatic buffers preserve surface protein integrity. However, viability with non-enzymatic methods may be lower (68.7% versus 93.2% for trypsin) [16], requiring potential yield trade-offs.
Neuronal Stem Cell Research and Differentiation
In neuronal stem cell applications, detachment method significantly impacts differentiation potential and stemness maintenance. Thermoresponsive surfaces show particular promise, as they enable non-invasive cell sheet harvest while preserving extracellular matrix components and cell-cell junctions [47]. This approach maintains the native microenvironment crucial for stem cell niche function. For passaging neuronal stem cells prior to differentiation, Accutase provides a balance between efficiency and preservation of differentiation capacity, though recovery periods may be necessary for surface marker reexpression [17].
High-Throughput Screening and Drug Development
For neuronal drug screening applications, emerging technologies offer significant advantages. Electrochemical detachment platforms achieve 95% detachment efficiency with >90% viability, enabling automated, reproducible processing compatible with high-throughput systems [9]. Similarly, lens-free imaging monitoring provides real-time, quantitative detachment assessment without manual intervention, standardizing this critical step across experimental batches [21]. These approaches enhance data consistency in screening campaigns evaluating neuroprotective compounds or neurotoxicity.
Cell Therapy and Regenerative Medicine Applications
In therapeutic neuronal cell preparation, detachment method selection must consider both regulatory and functional requirements. Traditional enzymatic methods introduce animal-derived components (trypsin) requiring rigorous validation and clearance [8]. Non-enzymatic approaches or defined recombinant enzymes (TrypLE) reduce regulatory burdens. For advanced applications like neuronal progenitor transplantation, thermoresponsive surfaces or electrochemical methods maintain critical surface properties and functionality essential for engraftment success [9] [47].
Emerging Technologies and Future Directions
The field of cell detachment is evolving with several promising technologies addressing limitations of current methods:
Nanostructured Stimuli-Responsive Surfaces: Advanced materials with decoupled adhesive and disjoining domains enable independent optimization of attachment and detachment functions [47]. These surfaces combine cell-adhesive epoxy photoresist domains with thermoresponsive PNIPAM brush domains, providing precise control over detachment initiation and progression.
AI-Integrated Monitoring and Control: Machine learning algorithms analyze detachment progression using lens-free imaging or other sensor data, enabling real-time process optimization [21] [48]. These systems automatically identify optimal inhibition points, improving reproducibility and cell quality.
Microfluidic Detachment Platforms: Systems integrating microfluidics, real-time imaging, and computational analysis enable gentle, targeted cell detachment using controlled hydrodynamic forces [48]. These platforms detect pre-detachment phases, minimizing mechanical stress and preserving cell integrity.
These innovations align with market trends toward automation and standardization, with the cell dissociation market projected to grow at 13.5-13.58% CAGR through 2035 [45] [49]. Pharmaceutical and biotechnology companies, representing 71.6% of market share, are driving adoption of these advanced technologies to improve reliability in cell-based therapeutic production [45].
Data-Driven Decisions: A Side-by-Side Analysis of Detachment Outcomes
The dissociation of tissues and cells into high-quality, viable single-cell suspensions is a critical first step in neuroscience research, enabling the study of neuronal function, development, and pathology. The choice of detachment method significantly impacts key performance metrics—cell viability, yield, and processing speed—which in turn dictate the success of downstream applications such as single-cell sequencing, electrophysiology, and long-term culture. This guide provides a quantitative comparison of enzymatic and non-enzymatic detachment methods, framing the analysis within the broader thesis that non-enzymatic approaches offer distinct advantages for preserving neuronal integrity and function while addressing the limitations of traditional enzymatic protocols.
The following analysis synthesizes experimental data from recent peer-reviewed literature and commercial protocols to offer researchers, scientists, and drug development professionals an evidence-based resource for selecting optimal detachment methodologies in neuronal research.
Comparative Performance Data
The table below summarizes quantitative data on viability, yield, and processing speed for various enzymatic and non-enzymatic detachment methods as reported in recent studies.
Table 1: Quantitative Performance Comparison of Cell Detachment Methods
| Method | Cell/Tissue Type | Viability (%) | Yield/ Efficiency | Processing Time | Key Advantages |
|---|---|---|---|---|---|
| Electrochemical [9] | Human cancer cells (osteosarcoma, ovarian) | >90% | 95% detachment efficiency | Minutes (specific duration not stated) | Minimal membrane damage; Animal-derived reagent-free |
| Hypersonic Levitation & Spinning (HLS) [18] | Human renal cancer tissue | 92.3% | 90% tissue utilization | 15 minutes | Excellent rare cell preservation; Non-contact |
| Optimized Enzymatic (Primary Neuron Kit) [50] | Mouse cortical neurons | 94-96% | ~4.5 x 10⁶ cells/mL (Mouse Cortex) | Protocol-dependent (hours) | High cell functionality; Established protocol |
| Traditional Trypsin [50] | Mouse cortical neurons | 83-92% | ~2.25 x 10⁶ cells/mL (Mouse Cortex) | Protocol-dependent (hours) | Widely available; Familiar methodology |
| Thermoresponsive Microcarriers (BrushGel) [51] | Human dermal fibroblasts, MSCs | >95% post-detachment | 65-69% detachment efficiency | Includes temperature shift | Enzyme-minimized; Scalable for biomanufacturing |
| Ultrasound Dissociation [12] | Bovine liver tissue, MDA-MB-231 cells | 91-98% | 53% ± 8% (sonication alone) | 30 minutes | Enzyme-free; Rapid processing |
Analysis of Methodologies
Non-Enzymatic Methods
Electrochemical Detachment
This novel approach applies low-frequency alternating voltage on a conductive biocompatible polymer surface to disrupt cell adhesion. The method achieves high detachment efficiency (95%) while maintaining excellent viability (>90%), overcoming limitations of enzymatic methods that can damage delicate cell membranes. The technique is particularly valuable for scalable biomanufacturing as it enables automated, contamination-conscious workflows and eliminates animal-derived reagents, which is crucial for therapeutic applications like CAR-T cell production [9].
Acoustic Methods (HLS and Ultrasound)
Hypersonic Levitation and Spinning (HLS) represents a breakthrough in non-contact tissue dissociation. Using a triple-acoustic resonator probe, this method generates microscale 'liquid jets' that exert precise hydrodynamic forces on tissues, causing them to levitate and execute a 'press-and-rotate' operation. This unique mechanism achieves 92.3% viability and 90% tissue utilization in just 15 minutes while better preserving rare cell populations compared to conventional methods [18].
Standard ultrasound dissociation techniques also show promise, achieving 91-98% viability with a 30-minute processing time, though with more moderate yield (53% with sonication alone) [12]. Both acoustic methods offer the significant advantage of being non-contact, minimizing mechanical stress on cells.
Thermoresponsive Microcarriers
The BrushGel platform utilizes gelatin methacryloyl (GelMA) hydrogel microcarriers coated with poly(N-isopropyl acrylamide) (PNIPAM) polymer brushes. This innovative material enables cell detachment through a simple temperature reduction from 37°C to 20°C, achieving >95% post-detachment viability with 65-69% detachment efficiency. While detachment efficiency is moderate, this approach reduces enzyme use by 10-fold, significantly lowering costs and avoiding enzyme-induced damage to cell membranes and surface proteins [51].
Enzymatic Methods
Optimized Enzymatic Protocol
Commercial optimized enzyme formulations (e.g., Thermo Scientific Pierce Primary Neuron Isolation Kit) demonstrate marked improvements over traditional trypsin, achieving 94-96% viability and approximately 2-fold higher cell yields from mouse embryonic cortical tissue. These optimized cocktails are specifically designed for sensitive neuronal cells, preserving neuronal functionality as evidenced by more intricate dendritic arbors and stronger synaptic protein expression compared to trypsin-based methods [50].
Traditional Trypsin
The conventional trypsin-based approach shows more variable performance, with viability ranging from 83-92% and substantially lower cell yields compared to optimized enzymatic methods. Trypsin's aggressive proteolytic activity can damage cell surface receptors and adhesion molecules, potentially compromising neuronal function in downstream applications [50].
Experimental Protocols
Non-Enzymatic Workflow
The following diagram illustrates the decision pathway for selecting and implementing non-enzymatic detachment methods:
Non-Enzymatic Method Selection Workflow
Electrochemical Detachment Protocol
- Principle: Application of alternating electrochemical current on a conductive biocompatible polymer nanocomposite surface to disrupt cell adhesion [9].
- Procedure:
- Culture cells on specialized conductive polymer nanocomposite surfaces.
- Apply low-frequency alternating voltage in a biocompatible buffer solution.
- Monitor detachment progress (typically completes within minutes).
- Collect released cells by gentle pipetting.
- Optimization Notes: Optimal frequency is critical for efficiency; identified ideal frequency increased detachment efficiency from 1% to 95% for human cancer cells [9].
Hypersonic Levitation and Spinning (HLS) Protocol
- Principle: Use of GHz-frequency acoustic waves to generate hypersonic streaming jets that levitate and spin tissue samples, applying precise hydrodynamic shear forces for dissociation [18].
- Procedure:
- Place tissue sample (e.g., human renal cancer tissue) in dissociation chamber filled with appropriate buffer.
- Activate triple-acoustic resonator probe to generate hypersonic streaming field.
- Allow tissue to levitate and undergo 'press-and-rotate' operation for 15 minutes.
- Collect dissociated cells from outlet chamber while debris is directed to waste chamber.
- Optimization Notes: Integrated apparatus automatically handles dissociation, fluid replacement, filtration, and output functions [18].
Enzymatic Workflow
The following diagram illustrates the optimized enzymatic protocol for neuronal cell isolation:
Optimized Enzymatic Dissociation Workflow for Neurons
Optimized Enzymatic Dissociation for Primary Neurons
- Principle: Controlled use of proteases to digest intercellular protein junctions followed by gentle mechanical disruption to liberate individual cells [50].
- Procedure:
- Rapidly dissect brain regions (cortex, hippocampus) from E17-18 rat embryos or E15 spinal cord in ice-cold HBSS [33].
- Mince tissue into 1-2 mm³ pieces using microdissection scissors.
- Digest with optimized enzyme formulation (e.g., Primary Neuron Isolation Kit) for 30-45 minutes at 37°C with gentle agitation.
- Triturate digested tissue 3-5 times through a fire-polished Pasteur pipette.
- Centrifuge cell suspension at 70-90 × g for 5 minutes.
- Resuspend pellet in neuronal culture medium (Neurobasal plus medium with B-27 supplement) [33].
- Plate cells on poly-D-lysine coated surfaces at desired density.
- Optimization Notes: Limit total dissection time to less than 1 hour; complete meninges removal is crucial for neuronal purity; enzymatic concentration and timing must be carefully optimized for specific tissue type [33].
The Scientist's Toolkit
Table 2: Essential Reagents and Materials for Neuronal Cell Dissociation
| Item | Function | Application Notes |
|---|---|---|
| Poly-D-Lysine (PDL) | Coats culture surfaces to enhance neuronal attachment | Critical for neuronal viability; excess can be toxic - ensure thorough rinsing [52] |
| Hanks' Balanced Salt Solution (HBSS) | Isotonic buffer for tissue dissection and washing | Maintained ice-cold to preserve tissue viability during dissection [33] |
| Primary Neuron Isolation Kit | Optimized enzyme formulation for neuronal tissue | Provides 2-fold higher yield vs. trypsin with 94-96% viability [50] |
| Neurobasal Medium with B-27 Supplement | Serum-free neuronal culture medium | Supports long-term neuronal health and reduces glial proliferation [33] |
| Trypan Blue | Viability stain for cell counting | Used with hemocytometer or automated cell counter for viability assessment [50] |
| Conductive Polymer Nanocomposite Surfaces | Specialized surfaces for electrochemical detachment | Enables enzyme-free detachment with >90% viability [9] |
| Thermoresponsive Microcarriers (BrushGel) | GelMA-based carriers with PNIPAM coating | Enables temperature-induced cell detachment with >95% viability [51] |
The quantitative comparison presented in this guide demonstrates that method selection involves balancing multiple performance metrics. Non-enzymatic methods (electrochemical, acoustic, thermoresponsive) generally provide superior cell viability and are preferred for sensitive applications where preserving native cell state is paramount. Optimized enzymatic approaches remain valuable for achieving high cell yields and are continually improving to minimize damage.
The evolving toolkit for cell detachment offers researchers multiple pathways to obtain high-quality neuronal preparations. Selection should be guided by the specific requirements of downstream applications, with non-enzymatic methods particularly advantageous for single-cell analyses, therapeutic manufacturing, and studies requiring maximal preservation of surface markers and functional integrity.
The investigation of neuronal phenotypes is fundamental to advancing our understanding of neuroscience and developing treatments for neurological disorders. A critical, yet often overlooked, step in these research workflows is the detachment of adherent neuronal cells from culture surfaces. The method chosen for cell detachment can significantly influence experimental outcomes by altering cell surface markers, viability, and functionality [8] [17].
This guide provides a comparative analysis of enzymatic and non-enzymatic cell detachment methods, focusing on their impact on neuronal phenotype. We objectively evaluate product performance by summarizing experimental data on key parameters including cell viability, surface marker integrity, and post-detachment functionality, providing researchers with evidence-based criteria for selecting the most appropriate detachment strategy.
Comparative Analysis of Detachment Methods
Cell detachment strategies primarily fall into two categories: enzymatic and non-enzymatic. Each method operates through a distinct mechanism and presents a unique profile of advantages and disadvantages for neuronal research.
Table 1: Overview of Cell Detachment Method Types, Mechanisms, and General Characteristics
| Method Type | Specific Examples | Mechanism of Action | Key Advantages | Key Disadvantages |
|---|---|---|---|---|
| Enzymatic | Trypsin, Accutase | Proteolytic cleavage of cell-adhesion proteins [8] [17]. | Rapid and effective for strongly adherent cells [16]. | Degrades surface proteins (e.g., FasL, Fas receptor); can dysregulate protein expression and metabolic pathways [8] [17]. |
| Non-enzymatic (Chemical) | EDTA-based buffers, Chelate-free solutions | Chelates calcium/magnesium ions, disrupting integrin-mediated adhesion [8] [17]. | Gentler on surface protein structure; preferred for flow cytometry [17]. | Less effective for strongly adherent cells; can require prolonged incubation or mechanical assistance [16] [17]. |
| Non-enzymatic (Physical/Novel) | Electrochemical, Thermo-responsive, Scraping | Electrochemical: Alters ionic microenvironment [9] [22].Thermo-responsive: Changes surface polymer hydration [8].Scraping: Mechanical force. | Electrochemical: High efficiency, >90% viability, automation-friendly [9] [22].Scraping: Preserves surface markers best [17]. | Scraping: Can cause cell tearing and death [17].Thermo-responsive: Requires precise control, can be less robust [8]. |
The diagram below illustrates the core decision-making workflow for selecting a detachment method based on the experimental goals.
Experimental Data and Performance Comparison
Quantitative Comparison of Cell Viability and Detachment Efficiency
The performance of detachment methods can be quantitatively assessed through cell viability and detachment efficiency metrics.
Table 2: Quantitative Comparison of Detachment Method Performance on Various Cell Types
| Detachment Method | Cell Type Tested | Viability (%) | Detachment Efficiency (%) | Key Experimental Findings | Source |
|---|---|---|---|---|---|
| Trypsin | Mesenchymal Stem Cells (MSCs) | 93.2% ± 3.2 | N/R | Significantly higher viability vs. enzyme-free buffer; higher reattachment rate post-thawing. | [16] |
| Enzyme-Free Dissociation Buffer | Mesenchymal Stem Cells (MSCs) | 68.7% ± 5.0 | N/R | Significantly lower viability and reattachment rate vs. trypsin. | [16] |
| Accutase | Macrophages (RAW264.7) | N/R | N/R | Maintained high cell counts after 60- and 90-minute treatments vs. other solutions. | [17] |
| Electrochemical Method | Human Cancer Cells (Osteosarcoma, Ovarian) | > 90% | 95% (from a baseline of 1%) | High efficiency and viability achieved using low-frequency alternating voltage. | [9] [22] |
N/R: Not Reported in the cited study
Impact on Surface Marker Integrity and Recovery
Preserving the integrity of cell surface proteins is critical for immunophenotyping and functional assays. Different detachment methods variably impact these markers.
Table 3: Impact of Detachment Methods on Surface Marker Expression and Recovery
| Detachment Method | Impact on Surface Markers | Evidence | Recovery Time |
|---|---|---|---|
| Accutase | Significant decrease in surface FasL and Fas receptor; cleaves extracellular portion of FasL. Does not affect F4/80 marker. | Flow cytometry showed reduced MFI; Western blot confirmed cleavage. | ~20 hours for FasL/Fas recovery. |
| EDTA-based Buffer | Mild impact; better preservation of FasL and Fas receptor compared to Accutase. | Higher MFI for FasL/Fas vs. Accutase-treated groups. | N/R |
| Scraping | Best preservation of surface markers. | Highest levels of surface FasL preserved. | N/R |
| Trypsin | Degrades most cell surface proteins; potential dysregulation of various protein expression levels. | Well-documented in literature; cleaves after lysine/arginine residues. | N/R |
N/R: Not Reported in the cited study
The following diagram summarizes the key effects of different detachment methods on a neuronal cell and the subsequent consequences for research.
Detailed Experimental Protocols
To ensure reproducibility and provide a clear framework for researchers, we outline the key methodologies from the cited experiments.
Protocol: Assessing Surface Marker Integrity Post-Detachment
This protocol is adapted from studies investigating the impact of accutase on surface proteins [17].
- Cell Culture: Grow adherent cells (e.g., RAW264.7 macrophages) to confluency in 12-well plates.
- Detachment:
- Treat cells with pre-warmed accutase or an EDTA-based dissociation buffer.
- Incubate at 37°C for 10-30 minutes, with gentle pipetting every 2-3 minutes to aid detachment.
- Flow Cytometry Analysis:
- Centrifuge the dissociated cell suspension and reconstitute the pellet in PBS.
- Follow standard staining procedures with antibodies against the surface markers of interest (e.g., FasL, Fas receptor).
- Analyze using a flow cytometer and compare the Mean Fluorescence Intensity (MFI) between treatment groups.
- Recovery Assay: After detachment with accutase, re-culture cells in complete medium for up to 20 hours. Harvest cells at different time points (e.g., 2h, 20h) to assess the recovery of surface marker expression via flow cytometry.
- Western Blot Analysis: To confirm cleavage of surface proteins, collect supernatant and cell lysates after detachment. Use an antibody targeting the extracellular portion of the protein to detect cleaved fragments.
Protocol: Novel Electrochemical Detachment Method
This protocol summarizes the novel enzyme-free strategy presented by MIT researchers [9] [22].
- Surface Preparation: Culture cells on a conductive, biocompatible polymer nanocomposite surface.
- Detachment Trigger: Apply a low-frequency alternating voltage to the culture surface. The specific optimal frequency must be determined experimentally.
- Process Monitoring: Disruption of cell adhesion typically occurs within minutes.
- Cell Harvesting and Analysis:
- Collect the detached cell suspension.
- Assess detachment efficiency by counting released cells versus remaining adherent cells.
- Determine cell viability using a standard trypan blue exclusion assay or an automated cell counter.
The Scientist's Toolkit: Essential Research Reagents and Materials
Selecting the right reagents is fundamental to the success of detachment and subsequent assays.
Table 4: Key Reagent Solutions for Cell Detachment and Phenotypic Analysis
| Reagent/Material | Function in Research | Key Considerations |
|---|---|---|
| Trypsin-EDTA | Gold-standard enzymatic detachment. Effective for robust cells. | Can over-digest surface proteins; concentration and incubation time must be optimized. |
| Accutase | A gentler, enzymatic blend for detaching sensitive cells. | Can still cleave specific proteins (e.g., FasL); requires recovery time for surface marker studies. |
| EDTA-based Dissociation Buffer | Non-enzymatic, chemical detachment. Ideal for surface marker preservation. | May be ineffective for strongly adherent cells without mechanical disruption or long incubation. |
| Electrochemical Surface | Novel platform for enzyme-free, physical detachment. | Enables high viability and automation; requires specialized cultureware. |
| Conductive Polymer Nanocomposite | The active surface for the electrochemical detachment method. | Key to the mechanism of ionic microenvironment disruption. |
| Trypan Blue | A vital dye for assessing cell viability post-detachment. | Distinguishes live (unstained) from dead (blue) cells; used with automated or manual counting. |
| Antibodies (e.g., anti-FasL) | Critical for flow cytometry to quantify surface marker expression. | Specificity and titer must be validated for the cell type and application. |
The choice between enzymatic and non-enzymatic cell detachment methods is a significant determinant in the reliability of data derived from neuronal phenotype studies. Enzymatic methods, while efficient, carry a high risk of altering the very cellular characteristics researchers aim to study, namely surface markers and functional integrity. Non-enzymatic strategies, particularly novel approaches like electrochemical detachment, demonstrate a superior ability to preserve cell viability and surface proteins, which is crucial for high-fidelity research in neuroscience and drug development.
Researchers must align their detachment protocol with their experimental endpoints. For studies where surface marker integrity is paramount, non-enzymatic methods are strongly recommended. As the field moves toward larger-scale, automated cell manufacturing for therapies, adopting gentle, efficient, and reproducible detachment technologies will be essential for progressing our understanding and treatment of neurological diseases.
In cellular research, particularly in the study of neurons and other adherent cells, the process of detaching cells from culture surfaces is a fundamental but critical step. The choice of detachment method can significantly influence experimental outcomes by affecting cell viability, surface protein integrity, and subsequent cellular functions. This case study focuses on comparing two common approaches: Accutase, an enzymatic dissociation solution, and EDTA-based buffers, which are non-enzymatic and work via calcium chelation. Within the broader thesis of evaluating enzymatic versus non-enzymatic detachment methods for neuronal research, we objectively examine how these methods differentially impact the expression and function of key surface proteins, with a specific focus on the Fas ligand (FasL) and its receptor. Understanding these effects is crucial for researchers, scientists, and drug development professionals who rely on accurate cell surface marker analysis for their work in immunology, neurobiology, and therapeutic development.
The following table summarizes the core comparative findings between Accutase and EDTA-based detachment methods regarding their effect on Fas ligand and Fas receptor expression.
| Parameter | Accutase | EDTA-Based Methods | Scraping (Control Reference) |
|---|---|---|---|
| Effect on Surface FasL | Significant decrease in Mean Fluorescence Intensity (MFI) [53] | Minimal decrease compared to scraping [53] | Preserved (highest level) [53] |
| Effect on Surface Fas Receptor | Significant decrease in MFI [53] | Preserved expression [53] | Information not specified in search results |
| Effect on Other Markers (e.g., F4/80) | No significant change [53] | No significant change [53] | Information not specified in search results |
| Mechanism of Action | Proteolytic cleavage; cleaves extracellular region of FasL [53] | Calcium chelation, disrupting cell adhesion [53] | Mechanical force [53] |
| Cell Recovery & Viability | High cell viability post-detachment [53] [46] | Lower cell viability compared to enzymatic methods [46] | Can cause cell damage [53] |
| Recovery Time for Surface Proteins | ~20 hours for FasL/Fas recovery [53] | Presumed immediate (not required) | Not applicable |
Detailed Experimental Data and Quantitative Comparisons
Impact on Specific Surface Protein Expression
Beyond the summary, a detailed analysis of flow cytometry data reveals the magnitude of the effects. Treatment with Accutase led to a significant reduction in the mean fluorescence intensity (MFI) of surface FasL on macrophages compared to cells treated with EDTA-based solutions (p < 0.001) [53]. The same detrimental effect was observed for the Fas receptor. Importantly, this was not a global effect on all surface proteins, as the expression of the macrophage-specific marker F4/80 remained unaltered by Accutase treatment, highlighting the specific vulnerability of FasL and Fas [53].
The duration of exposure during detachment also plays a role. While a 10-minute Accutase treatment significantly reduced surface FasL levels, extending the incubation to 30 minutes did not cause a further significant decrease in the context of that particular experiment [53]. This suggests that the proteolytic action occurs rapidly upon exposure.
Cell Viability and Recovery Post-Detachment
A key advantage of enzymatic methods is their efficiency and gentleness on overall cell health. Studies confirm that Accutase yields highly efficient recovery of viable cells [46]. Viability assays demonstrated that viable cell counts were significantly higher in Accutase-treated groups even after extended incubation periods (60-90 minutes) compared to EDTA-based solutions or DPBS buffers (p < 0.01) [53].
However, the compromised surface proteins require a recovery period. After Accutase treatment, cells need to be incubated in a complete culture medium for up to 20 hours for the surface levels of FasL and Fas receptor to return to normal [53]. This recovery period is a critical consideration for planning subsequent functional assays.
Experimental Protocols for Key Methodologies
Protocol: Comparing Detachment Methods for Flow Cytometry
The following detailed protocol is adapted from the methodologies used in the cited studies to compare the effects of Accutase and EDTA on surface marker expression [53] [46].
- Cell Culture and Preparation: Grow adherent cells (e.g., RAW264.7 macrophages or other relevant cell lines) to ~80% confluence in standard culture plates.
- Detachment:
- Accutase Group: Remove culture medium and wash cells with phosphate-buffered saline (PBS) without Ca2+/Mg2+. Add pre-warmed Accutase solution to cover the cell layer. Incubate at 37°C for 5-10 minutes, or until cells detach. Monitor under a microscope to avoid over-digestion.
- EDTA-Based Group: Wash cells similarly. Add a pre-warmed EDTA-based solution (e.g., Versene or PBS-based cell dissociation buffer with 1-5 mM EDTA). Incubate at 37°C for approximately 15-20 minutes, possibly with gentle scraping or pipetting to aid detachment of strongly adherent cells [53] [16].
- Scraping Group (Control): Use a cell scraper to mechanically dislodge cells in PBS on ice. This method helps establish a baseline for surface protein preservation [53].
- Neutralization and Collection: Neutralize the Accutase or EDTA solution with complete culture medium containing serum. Gently pipette the cells, transfer the suspension to a tube, and centrifuge to pellet the cells.
- Flow Cytometry Staining: Resuspend the cell pellet in flow cytometry staining buffer. Aliquot cells for different antibody panels. Stain with fluorescently-labeled antibodies against FasL (CD178), Fas (CD95), and a stable control marker (e.g., F4/80 for macrophages). Include appropriate isotype controls. Analyze cells on a flow cytometer, gating on live cells based on viability dye exclusion [53] [54].
- Recovery Assay: For Accutase-treated cells, re-seed a portion into a new culture dish with complete medium and analyze FasL/Fas expression via flow cytometry after 2, 6, and 20 hours to monitor protein recovery [53].
Protocol: Assessing Functional Consequences
To determine if the loss of surface proteins translates to functional deficits, a FasL-mediated apoptosis assay can be performed.
- Detach and Recover: Detach effector cells (e.g., FasL-expressing lymphocytes) using Accutase and EDTA as described. Allow a subpopulation of Accutase-treated cells to recover for 20 hours [53].
- Co-culture: Co-culture the treated effector cells with target cells that express the Fas receptor (e.g., certain lymphocyte cell lines) in a standard co-culture plate.
- Measure Apoptosis: After an appropriate co-culture period (e.g., 6-24 hours), measure apoptosis in the target cell population. This can be done using:
- Flow cytometry with Annexin V/propidium iodide (PI) staining.
- Caspase activity assays (e.g., Caspase-3/7 glow assay).
- Analysis: Compare the level of target cell apoptosis induced by effector cells detached with different methods. A reduced apoptotic rate in the Accutase group (without recovery) would indicate a functional impairment due to FasL cleavage.
Mechanisms and Signaling Pathways
The fundamental mechanism behind the differential effects of Accutase and EDTA lies in their mode of action. EDTA is a calcium chelator that disrupts calcium-dependent integrins, which are essential for cell-to-substrate and cell-to-cell adhesion. This is a physical disruption that generally leaves surface proteins intact [53]. In contrast, Accutase is a proteolytic enzyme mixture that actively cleaves peptide bonds in proteins that anchor the cell to the substrate.
Diagram 1: Mechanism of Action for Cell Detachment Methods. EDTA chelates calcium, disrupting integrin-mediated adhesion without damaging most surface proteins. Accutase cleaves both adhesion proteins and specific surface markers like FasL, releasing soluble fragments.
As illustrated in Diagram 1, Accutase's enzymatic activity is not limited to adhesion molecules. Western blot analysis has confirmed that Accutase cleaves the extracellular region of FasL into small fragments under 20 kD, which are then detected in the cell supernatant. This cleavage explains the loss of membrane-bound FasL and the consequent reduction in fluorescence signal observed in flow cytometry [53]. Immunofluorescence staining further supports this, showing that after Accutase treatment, FasL is largely absent from the cell membrane, whereas it remains membrane-bound in EDTA-treated cells [53].
The Scientist's Toolkit: Essential Research Reagents
The table below lists key reagents and their functions used in the experiments cited in this guide, providing a practical resource for researchers.
| Reagent / Material | Function / Application | Key Consideration / Effect |
|---|---|---|
| Accutase [53] [55] | Enzymatic cell detachment solution for dissociating adherent cells. | Gentle on viability but cleaves specific surface proteins (e.g., FasL, Fas). |
| EDTA-Based Dissociation Buffer (e.g., Versene) [53] [16] | Non-enzymatic, calcium-chelating solution for cell detachment. | Preserves surface protein integrity but may be less effective for strongly adherent cells. |
| Flow Cytometer with Antibodies (anti-FasL, anti-Fas, viability dye) [53] [54] | Quantifying surface protein expression (MFI) and cell viability post-detachment. | Essential for objectively comparing the impact of different detachment methods. |
| Cell Culture Plates with Coating (e.g., extracellular matrix) [56] | Providing a surface for adherent cell growth (neurons, macrophages, etc.). | Coating type can influence adhesion strength and required detachment force. |
| Complete Cell Culture Medium (with serum) [53] | Neutralizing enzymatic activity after detachment and supporting cell recovery. | Critical for stopping the reaction and allowing surface protein re-synthesis. |
| Western Blot Equipment & Antibodies [53] | Detecting cleavage of surface proteins (e.g., FasL fragments in supernatant). | Provides mechanistic evidence for proteolytic activity beyond flow cytometry. |
This case study demonstrates that the choice between Accutase and EDTA-based detachment methods involves a direct trade-off between cell viability and the preservation of specific surface epitopes. While Accutase offers excellent cell recovery and is gentler on overall cell health, it significantly compromises the expression of FasL and Fas receptor through proteolytic cleavage. EDTA-based methods, though potentially harsher on viability and less effective for some strongly adherent cells, are superior for preserving the native structure of these and potentially other sensitive surface proteins.
Based on the evidence, we recommend the following for researchers:
- Use EDTA-based methods or scraping when studying FasL/Fas receptor expression or function, or when analyzing other surface markers known to be sensitive to proteolysis (e.g., CD163, CD206) [53] [46].
- Accutase is a suitable choice for experiments where high viability is paramount and the surface proteins of interest are known to be resistant to its activity (e.g., CD14, CD117) [53], or for applications like subculturing where protein integrity is not immediately critical.
- Incorporate a recovery period: If Accutase must be used, allow cells to recover in culture for at least 20 hours before assaying FasL/Fas-dependent functions [53].
- Validate for your specific system: The impact of detachment can vary by cell type. Researchers should empirically compare methods using flow cytometry and functional assays for their specific models to ensure methodological rigor and reproducible results.
Scalability and Contamination Risk Assessment for Biomanufacturing Workflows
In the field of biomanufacturing, particularly for cell-based therapies and regenerative medicine, the process of detaching adherent cells from culture surfaces represents a critical step with profound implications for both scalability and contamination risk. The choice between enzymatic and non-enzymatic detachment methods directly influences cell viability, functionality, and regulatory compliance, making it a pivotal consideration for researchers and drug development professionals. This guide provides an objective comparison of these technologies, focusing on their performance in scalable biomanufacturing workflows and their associated contamination profiles, with specific consideration for applications in neuroscience research.
The global cell dissociation market, valued at approximately USD 406 million in 2024 and projected to reach USD 1.62 billion by 2035, reflects the growing importance of these technologies in the biotechnology and pharmaceutical sectors [45] [57]. This growth is largely driven by increasing investments in cell-based therapies and the need for more efficient, scalable cell processing methods.
Fundamental Mechanisms
Enzymatic methods primarily utilize proteolytic enzymes such as trypsin, collagenase, or accutase to cleave adhesion proteins and extracellular matrix components that anchor cells to culture surfaces [8] [17]. These methods are effective but inherently invasive, as they target and degrade cell surface proteins during the detachment process.
Non-enzymatic methods employ alternative approaches including:
- Chelating agents (e.g., EDTA) that bind calcium and magnesium ions essential for integrin-mediated adhesion [17]
- Electrochemical approaches that use alternating current on conductive surfaces to disrupt cell adhesion [9] [22]
- Thermo-responsive polymers that change properties with temperature shifts to release cells [58]
- Physical methods including mechanical scraping, magnetic fields, or light-based detachment [8]
Experimental Protocols for Method Evaluation
To objectively compare detachment methods, researchers employ standardized experimental protocols:
Cell Viability Assessment:
- Dissociate confluent monolayers with either enzymatic or non-enzymatic agents
- Centrifuge cell suspension and discard supernatant
- Reconstitute cell pellet in phosphate-buffered saline (PBS)
- Analyze cell viability using trypan blue exclusion assay with an automated cell counter [16]
Re-attachment Efficiency Testing:
- Seed dissociated cells onto fresh culture dishes
- After 24 hours of culture, wash off unattached cells
- Subject reattached cells to MTT assay: add 1 mg/ml MTT solution to each well
- Incubate for 3 hours at 37°C in the dark
- Remove MTT solution and wash cells with PBS
- Extract MTT-formazan products with DMSO
- Measure absorbance spectrophotometrically at 570 nm [16]
Surface Protein Integrity Analysis:
- Treat cells with detachment solutions according to manufacturer instructions
- Harvest cells and stain with antibodies against specific surface markers (e.g., FasL, Fas receptor)
- Analyze by flow cytometry to measure mean fluorescence intensity
- For immunofluorescence, fix cells and stain with target antibodies and F-actin markers
- Image using confocal microscopy to determine membrane localization of surface proteins [17]
Comparative Performance Analysis
Quantitative Comparison of Detachment Methods
Table 1: Direct Performance Comparison of Cell Detachment Methods
| Parameter | Enzymatic Methods | Non-Enzymatic Chemical Methods | Electrochemical Method (MIT) | Physical/Scraping |
|---|---|---|---|---|
| Cell Viability | 93.2% (trypsin on MSC) [16] | 68.7% (buffer on MSC) [16] | >90% [9] [22] | Variable, risk of damage |
| Detachment Efficiency | High (95%+) | Moderate | 95% [9] | Complete but destructive |
| Surface Protein Damage | Significant degradation of membrane proteins [8] [17] | Minimal when optimized | Preserved (>90% viability) [22] | Physical disruption |
| Recovery Time Post-Detachment | Required for surface protein regeneration | Shorter | Not specified | Extensive for membrane repair |
| Scalability | Well-established but limited by enzyme costs and residuals | Moderate, limited by effectiveness on complex tissues | High potential for uniform application across large areas [22] | Low, manual intensive |
| Therapeutic Compatibility | Limited by animal-derived components and enzyme residuals | Higher, especially with defined chemical formulations | High, no enzyme contaminants [9] | Low, inconsistent results |
Scalability Assessment for Biomanufacturing
Table 2: Scalability and Manufacturing Considerations
| Factor | Enzymatic Dissociation | Non-Enzymatic Dissociation |
|---|---|---|
| Current Market Share | 47.9% (dominant position) [45] | Growing segment, predicted fastest CAGR [57] |
| Process Integration | Compatible with existing workflows but requires removal steps | Easier integration for closed-loop systems [22] |
| Automation Potential | High but complicated by enzyme inactivation steps | Higher, enabling fully automated systems [9] |
| Volume Capability | Limited by enzyme costs and availability | More scalable with synthetic materials |
| Batch Consistency | Variable due to enzyme lot differences | Potentially higher with defined processes [8] |
| Regulatory Pathway | Established but complicated by animal-derived components | Streamlined with defined, synthetic components [9] |
Contamination Risk Profile
Table 3: Contamination and Risk Assessment
| Risk Category | Enzymatic Methods | Non-Enzymatic Methods |
|---|---|---|
| Introduction of Animal-Derived Components | High (trypsin often sourced from animals) [9] | None with synthetic formulations [57] |
| Process-Related Impurities | Enzyme residuals requiring removal [8] | Chemical residuals potentially easier to remove |
| Impact on Final Product Purity | Significant concern for therapies | Reduced concern |
| Microbial Contamination Risk | Standard | Standard |
| Cross-Contamination in Reuse | Higher with enzyme carryover | Lower with disposable systems |
| Supply Chain Vulnerabilities | Medium-High (enzyme supply disruptions) [45] | Medium (specialized materials) |
Technological Innovations and Emerging Solutions
Advanced Non-Enzymatic Platforms
Recent research has yielded promising alternatives to conventional detachment methods:
Electrochemical Cell Detachment (MIT Platform):
- Utilizes alternating electrochemical current on conductive biocompatible polymer nanocomposite surfaces
- Achieves 95% detachment efficiency with >90% cell viability
- Operates by applying low-frequency alternating voltage to disrupt adhesion
- Enables detachment within minutes without enzymatic treatment [9] [22]
Thermo-Responsive Coatings (NSF-Funded Research):
- Implements polymer coatings with temperature-dependent properties
- Allows cell adhesion and proliferation at standard culture temperatures
- Triggers cell detachment when temperature is reduced
- Eliminates enzyme contamination while maintaining cell integrity [58]
Signaling Pathways in Cell Detachment
The following diagram illustrates the fundamental mechanisms of cell adhesion and how different detachment methods interact with these pathways:
Cell Adhesion and Detachment Mechanisms: This diagram illustrates how cells adhere to surfaces through extracellular matrix (ECM) interactions and how different detachment methods target these adhesion mechanisms. Enzymatic methods directly cleave proteins, while non-enzymatic approaches use alternative disruption mechanisms.
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Materials for Cell Detachment Research
| Reagent/Equipment | Function | Application Context |
|---|---|---|
| Trypsin-EDTA | Proteolytic enzyme chelating agent combination | Standard enzymatic dissociation for robust cell types [16] |
| Accutase | Mild enzymatic blend of proteolytic and collagenolytic enzymes | Gentle dissociation for sensitive cells; but may cleave specific surface proteins [17] |
| EDTA-Based Dissociation Buffer | Calcium and magnesium chelation disrupting integrin binding | Non-enzymatic dissociation preserving surface proteins [17] |
| Thermo-Responsive Polymers | Temperature-sensitive coatings that release cells upon cooling | Non-enzymatic harvesting with minimal cellular damage [58] |
| Conductive Polymer Nanocomposite | Electrochemically active surface for current-induced detachment | Enzyme-free, high-efficiency detachment for scalable bioprocessing [9] [22] |
| Automated Cell Counter | Viability assessment via trypan blue exclusion | Standardized evaluation of detachment method efficacy [16] |
| MTT Assay Components | Tetrazolium dye for metabolic activity measurement | Post-detachment cell functionality and reattachment capacity [16] |
Implications for Neuronal Research
The detachment method selection carries particular significance in neuronal research, where preservation of surface receptors and maintenance of functional integrity are paramount. Neuronal cells often exhibit heightened sensitivity to enzymatic treatment, making non-enzymatic approaches potentially advantageous:
- Surface Receptor Preservation: Non-enzymatic methods better maintain neurotransmitter receptors and signaling complexes essential for neuronal function [17]
- Neurite Integrity: Gentle detachment minimizes damage to delicate neurite processes
- Electrical Properties: Electrochemical methods may offer compatibility with electrophysiological studies
- Differentiation Maintenance: Reduced disruption of surface markers important for neuronal differentiation status
The assessment of scalability and contamination risks in biomanufacturing workflows reveals a shifting landscape where non-enzymatic detachment methods present compelling advantages for specific applications, particularly in sensitive fields like neuronal research. While enzymatic methods currently dominate the market and offer proven effectiveness for many cell types, emerging technologies in the non-enzymatic domain address critical limitations related to contamination risks, surface protein damage, and therapeutic compatibility.
For researchers and biomanufacturing professionals, selection criteria should extend beyond immediate efficiency to consider downstream applications, regulatory pathways, and final product quality. The continuing evolution of detachment technologies promises enhanced capabilities for scalable, contamination-conscious manufacturing of cell-based therapies, with non-enzymatic methods positioned for increasing adoption as validation data accumulates and implementation barriers are addressed.
Conclusion
The choice between enzymatic and non-enzymatic detachment is not one-size-fits-all but must be strategically aligned with experimental goals. While enzymatic methods offer speed and efficiency, they carry a significant risk of damaging critical neuronal surface proteins, with recovery times extending up to 20 hours. Non-enzymatic methods better preserve membrane integrity but may be less effective for strongly adherent cells and require mechanical assistance. The future of neuronal biomanufacturing lies in novel, gentle technologies like the electrochemical platform developed at MIT, which achieves over 90% viability and 95% efficiency without enzymes. For translational research in cell therapies, regenerative medicine, and high-throughput drug screening, adopting these advanced, controlled detachment methods will be crucial for generating reliable, reproducible, and clinically relevant neural models.
References
- 1. Extracellular Matrix Signaling Cues: Biological Functions ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12271642/]
- 2. Extracellular Matrix: Functions in the Nervous System - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC3003458/]
- 3. Measurement Systems for Cell Adhesive Forces - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC4347357/]
- 4. Co-administration of extracellular matrix-based ... [https://www.frontiersin.org/journals/neuroscience/articles/10.3389/fnins.2023.1177040/full]
- 5. Chitosan suitability to adhesion, neuronal differentiation ... [https://www.sciencedirect.com/science/article/pii/S2666893925000258]
- 6. The Role of Extracellular Matrix (ECM) Adhesion Motifs in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10302969/]
- 7. Region-specific brain decellularized extracellular matrix ... [https://www.nature.com/articles/s41598-025-95656-w]
- 8. Cell detachment: A review of techniques, challenges, and ... [https://www.sciencedirect.com/science/article/abs/pii/S0079610725000173]
- 9. An improved way to detach cells from culture surfaces [https://news.mit.edu/2025/improved-way-detach-cells-culture-surfaces-1118]
- 10. Methods for isolating highly-enriched embryonic spinal ... [https://www.sciencedirect.com/science/article/abs/pii/S0165027006002342]
- 11. Polyethyleneimine facilitates the growth and ... [https://www.nature.com/articles/s41598-024-77710-1]
- 12. Recent advancements in tissue dissociation techniques for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12622302/]
- 13. Microscaled Cell Surface Proteomics for Cryo-preserved ... [https://www.sciencedirect.com/science/article/pii/S1535947625005365]
- 14. A More Rapid Method for Culturing LUHMES-Derived ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12299948/]
- 15. Label-Free Long-Term Methods for Live Cell Imaging of ... [https://www.mdpi.com/2079-6374/13/3/404]
- 16. Comparison of Enzymatic and Non- ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3055293/]
- 17. Different methods of detaching adherent cells and their ... [https://www.nature.com/articles/s41598-022-09605-y]
- 18. Hypersonic levitation and spinning: paving the way for ... [https://www.nature.com/articles/s44172-025-00497-0]
- 19. The Importance of Tissue Dissociation Enzymes in Cell & ... [https://levitasbio.com/the-importance-of-tissue-dissociation-enzymes-in-cell-tissue-culture/]
- 20. Effect of enzymatic and mechanical methods of dissociation ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4785092/]
- 21. A novel feature for monitoring the enzymatic harvesting ... [https://www.nature.com/articles/s41598-022-22561-x]
- 22. Researchers present novel, enzyme-free strategy for ... [https://meche.mit.edu/news-media/researchers-present-novel-enzyme-free-strategy-detaching-cells-culture-surfaces]
- 23. A detailed comparison of common dissociation methods [https://www.sciencedirect.com/science/article/pii/S2468498820300093]
- 24. Recent advancements in tissue dissociation techniques for ... [https://pubmed.ncbi.nlm.nih.gov/41248140/]
- 25. Automated manufacturing of cell therapies [https://www.sciencedirect.com/science/article/pii/S0168365925001701]
- 26. Enzymatic Detachment of Therapeutic Mesenchymal ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3905350/]
- 27. Accutase™ - Capricorn's gentle Detachment Solution [https://www.capricorn-scientific.com/knowledge-center/accutase-trypsin-alternative]
- 28. A non-aggressive, highly efficient, enzymatic method for ... [https://bmcneurosci.biomedcentral.com/articles/10.1186/s12868-016-0262-y]
- 29. Proposed practical protocol for flow cytometry analysis of ... [https://www.frontiersin.org/journals/cellular-neuroscience/articles/10.3389/fncel.2022.1017976/full]
- 30. Enzymatic and Mechanical Approaches for Mammalian ... [https://orbitalshakers.net/blogs/blog/enzymatic-and-mechanical-approaches-for-mammalian-tissue-dissociation]
- 31. Manufacturing Cells for Clinical Use - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC4921138/]
- 32. of Chain-Forming Neuroblasts by Fyn-Mediated Control of... Detachment [https://pmc.ncbi.nlm.nih.gov/articles/PMC6705933/]
- 33. Optimized Protocols for the Isolation and Culture of ... [https://www.integrmed.org/journal/view.php?number=64]
- 34. A Sensory Neuron Subpopulation with Unique Sequential ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6762266/]
- 35. snCED-seq: high-fidelity cryogenic enzymatic dissociation ... [https://www.nature.com/articles/s41467-025-59464-0]
- 36. Cell Dissociation Market Size 2025 to 2034 [https://www.precedenceresearch.com/cell-dissociation-market]
- 37. Responsive systems for cell sheet detachment - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC3812292/]
- 38. Ultrafast light-induced charge and thermal transport effects in ... [https://www.the-innovation.org/article/doi/10.59717/j.xinn-mater.2025.100168]
- 39. The likely role of proteolytic enzymes in unwanted ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5137914/]
- 40. 神经细胞类型的流式细胞仪协议的表面和细胞内抗原分析 [https://www.jove.com/cn/t/52241/flow-cytometry-protocols-for-surface-intracellular-antigen-analyses]
- 41. Cell-Surface Marker Signatures for the Isolation of Neural ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3047583/]
- 42. of Comparison and Enzymatic - Non Means of Dissociating... Enzymatic [https://biologicalproceduresonline.biomedcentral.com/articles/10.1007/s12575-009-9001-4]
- 43. for isolating highly-enriched embryonic spinal cord Methods ... neurons [https://pubmed.ncbi.nlm.nih.gov/16787666/]
- 44. and Enzymatic - Non Cell... | Thermo Fisher Scientific - KR Enzymatic [https://www.thermofisher.com/kr/ko/home/references/gibco-cell-culture-basics/cell-culture-protocols/cell-dissociation-protocol.html]
- 45. Cell Dissociation Market Demand 2025-2035 [https://www.futuremarketinsights.com/reports/cell-dissociation-market]
- 46. Impact of Detachment Methods on M2 Macrophage ... [https://www.sciencedirect.com/science/article/abs/pii/S0022175915300284]
- 47. Strategy for Nonenzymatic Harvesting of Cells via ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10614186/]
- 48. Preliminary Study on Sensor-Based Detection of an Adherent ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12074402/]
- 49. Cell Dissociation Market Size Surges 13.58% CAGR by 2034 [https://www.towardshealthcare.com/insights/cell-dissociation-market-sizing]
- 50. An Efficient Method for the Isolation of Highly Functional ... [https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/protein-biology-application-notes/efficient-method-isolation-highly-functional-primary-neurons.html]
- 51. Thermoresponsive BrushGel Microcarriers for Efficient Cell ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12417781/]
- 52. Optimized protocol for obtaining and characterizing primary ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9558106/]
- 53. Different methods of detaching adherent cells and their effects on the... [https://www.nature.com/articles/s41598-022-09605-y?error=cookies_not_supported&code=b6d2c73c-bd46-44ae-a095-7a98a00165cf]
- 54. Substantial loss of T cells upon lymphocyte isolation from ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12436355/]
- 55. ™ Cell ACCUTASE Solution | STEMCELL Technologies Detachment [https://www.stemcell.com/products/accutase.html]
- 56. A Co- culture to Model Method -Oligodendrocyte Interactions Neuron [https://www.jove.com/t/31069/a-co-culture-method-to-model-neuron-oligodendrocyte-interactions]
- 57. Cell Dissociation Market Size Projected to Attain USD [https://www.globenewswire.com/news-release/2025/08/20/3136569/0/en/Cell-Dissociation-Market-Size-Projected-to-Attain-USD-1-451-89-Million-by-2034.html]
- 58. PFI-TT: Non-enzymatic harvesting of cell cultures [https://nsf.elsevierpure.com/en/projects/pfi-tnon-enzymatic-harvesting-of-cell-cultures/]